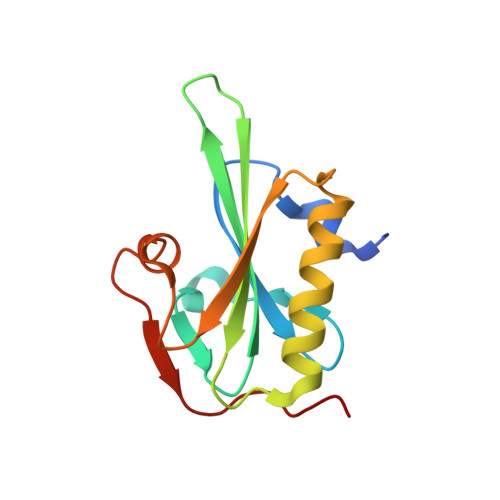
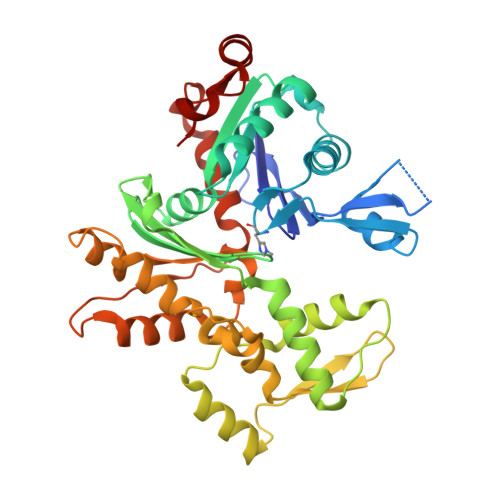
Structural basis for actin assembly, activation of ATP hydrolysis, and delayed phosphate release
Murakami, K., Yasunaga, T., Noguchi, T.Q.P., Gomibuchi, Y., Ngo, K.X., Uyeda, T.Q.P., Wakabayashi, T.(2010) Cell 143: 275-287
- PubMed: 20946985
- DOI: https://doi.org/10.1016/j.cell.2010.09.034
- Primary Citation of Related Structures:
3A5L, 3A5M, 3A5N, 3A5O, 3G37 - PubMed Abstract:
Assembled actin filaments support cellular signaling, intracellular trafficking, and cytokinesis. ATP hydrolysis triggered by actin assembly provides the structural cues for filament turnover in vivo. Here, we present the cryo-electron microscopic (cryo-EM) structure of filamentous actin (F-actin) in the presence of phosphate, with the visualization of some α-helical backbones and large side chains. A complete atomic model based on the EM map identified intermolecular interactions mediated by bound magnesium and phosphate ions. Comparison of the F-actin model with G-actin monomer crystal structures reveals a critical role for bending of the conserved proline-rich loop in triggering phosphate release following ATP hydrolysis. Crystal structures of G-actin show that mutations in this loop trap the catalytic site in two intermediate states of the ATPase cycle. The combined structural information allows us to propose a detailed molecular mechanism for the biochemical events, including actin polymerization and ATPase activation, critical for actin filament dynamics.
Organizational Affiliation:
Department of Biosciences, School of Science and Engineering, Teikyo University, Toyosatodai 1-1, Utsunomiya 320-8551, Japan.