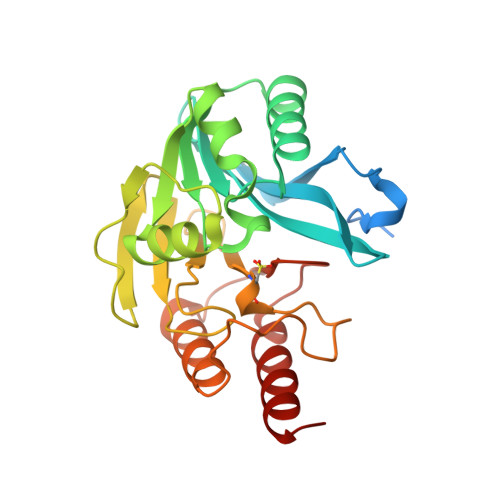
Discovery of Novel Inhibitor Scaffolds Against the Metallo-Beta-Lactamase Vim-2 by Spr Based Fragment Screening
Christopeit, T., Carlsen, T.J.O., Helland, R., Leiros, H.K.S.(2015) J Med Chem 58: 8671
- PubMed: 26477515
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01289
- Primary Citation of Related Structures:
5ACU, 5ACV, 5ACW, 5ACX - PubMed Abstract:
Metallo-β-lactamase (MBL) inhibitors can restore the function of carbapenem antibiotics and therefore help to treat infections of antibiotic resistant bacteria. In this study, we report novel fragments inhibiting the clinically relevant MBL Verona integron-encoded metallo-β-lactamase (VIM-2). The fragments were identified from a library of 490 fragments using an orthogonal screening approach based on a surface plasmon resonance (SPR) based assay combined with an enzyme inhibition assay. The identified fragments showed IC50 values between 14 and 1500 μM and ligand efficiencies (LE) between 0.48 and 0.23 kcal/mol per heavy atom. For two of the identified fragments, crystal structures in complex with VIM-2 were obtained. The identified fragments represent novel inhibitor scaffolds and are good starting points for the design of potent MBL inhibitors. Furthermore, the established SPR based assay and the screening approach can be adapted to other MBLs and in this way improve the drug discovery process for this important class of drug targets.
Organizational Affiliation:
Department of Chemistry, Faculty of Science and Technology, The Norwegian Structural Biology Centre (NorStruct), UiT The Arctic University of Norway , N-9037 Tromsø, Norway.