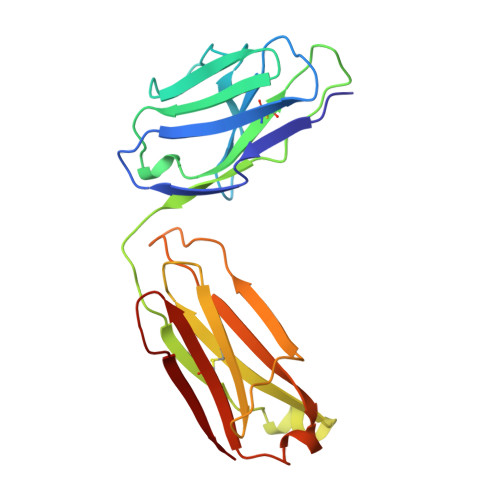
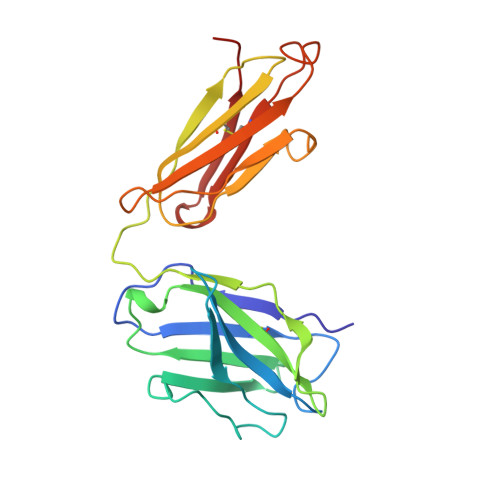
Structural studies on an inhibitory antibody against Thermus aquaticus DNA polymerase suggest mode of inhibition.
Murali, R., Helmer-Citterich, M., Sharkey, D.J., Scalice, E.R., Daiss, J.L., Sullivan, M.A., Krishna Murthy, H.M.(1998) Protein Eng 11: 79-86
- PubMed: 9605541
- DOI: https://doi.org/10.1093/protein/11.2.79
- Primary Citation of Related Structures:
1AY1 - PubMed Abstract:
TP7, an antibody against Thermus aquaticus DNA polymerase I (TaqP), is used as a thermolabile switch in 'hot start' variations of PCR to minimize non-specific amplification events. Earlier studies have established that TP7 binds to the polymerase domain of TaqP, competes with primer template complex for binding and is a potent inhibitor of the polymerase activity of TaqP. We report crystallographic determination of the structure of an Fab fragment of TP7 and computational docking of the structure with the known three-dimensional structure of the enzyme. Our observations strongly suggest that the origin of inhibitory ability of TP7 is its binding to enzyme residues involved in DNA binding and polymerization mechanism. Although criteria unbiased by extant biochemical data have been used in identification of a putative solution, the resulting complex offers an eminently plausible structural explanation of biochemical observations. The results presented are of general significance for interpretation of docking experiments and in design of small molecular inhibitors of TaqP, that are not structurally similar to substrates, for use in PCR. Structural and functional similarities noted among DNA polymerases, and the fact that several DNA polymerases are pharmacological targets, make discovery of non-substrate based inhibitors of additional importance.
Organizational Affiliation:
Department of Pathology and Laboratory Medicine, University of Pennsylvania, Philadelphia 19104, USA.