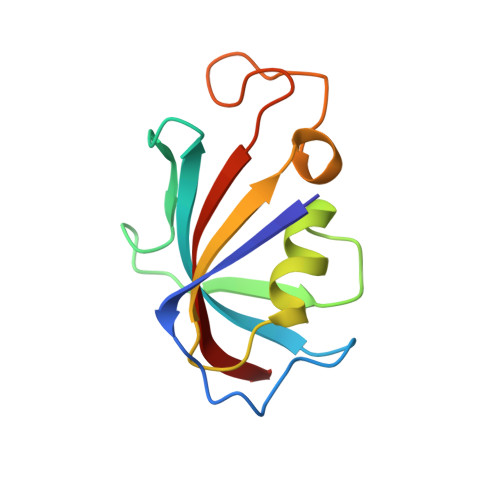
32-Indolyl ether derivatives of ascomycin: three-dimensional structures of complexes with FK506-binding protein.
Becker, J.W., Rotonda, J., Cryan, J.G., Martin, M., Parsons, W.H., Sinclair, P.J., Wiederrecht, G., Wong, F.(1999) J Med Chem 42: 2798-2804
- PubMed: 10425089
- DOI: https://doi.org/10.1021/jm9806042
- Primary Citation of Related Structures:
1QPF, 1QPL - PubMed Abstract:
32-Indole ether derivatives of tacrolimus and ascomycin retain the potent immunosuppressive activity of their parent compounds but display reduced toxicity. In addition, their complexes with the 12-kDa FK506-binding protein (FKBP) form more stable complexes with the protein phosphatase calcineurin, the molecular target of these drugs. We have solved the three-dimensional structures of the FKBP complexes with two 32-indolyl derivatives of ascomycin. The structures of the protein and the macrolide are remarkably similar to those seen in the complexes with tacrolimus and ascomycin. The indole groups project away from the body of the complex, and multiple conformations are observed for the linkage to these groups as well as for a nearby peptide suggesting apparent flexibility in these parts of the structure. Comparison of these structures with that of the ternary complex of calcineurin, FKBP, and tacrolimus suggests that the indole groups interact with a binding site comprising elements of both the calcineurin alpha- and beta-chains and that this interaction is responsible for the increased stability of these complexes.
Organizational Affiliation:
Departments of Endocrinology and Chemical Biology, Medicinal Chemistry, and Immunology Research, Merck Research Laboratories, P.O. Box 2000, Rahway, New Jersey 07065-0900, USA. beckerj@merck.com