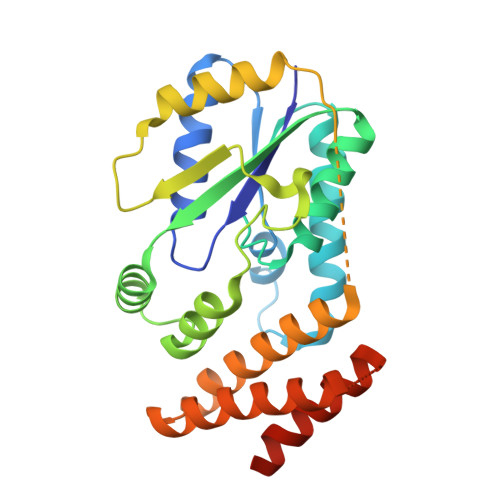
Structure of a tRNA-dependent kinase essential for selenocysteine decoding
Araiso, Y., Sherrer, R.L., Ishitani, R., Ho, J.M.L., Soll, D., Nureki, O.(2009) Proc Natl Acad Sci U S A 106: 16215-16220
- PubMed: 19805283
- DOI: https://doi.org/10.1073/pnas.0908861106
- Primary Citation of Related Structures:
3A4L, 3A4M, 3A4N - PubMed Abstract:
Compared to bacteria, archaea and eukaryotes employ an additional enzyme for the biosynthesis of selenocysteine (Sec), the 21(st) natural amino acid (aa). An essential RNA-dependent kinase, O-phosphoseryl-tRNA(Sec) kinase (PSTK), converts seryl-tRNA(Sec) to O-phosphoseryl-tRNA(Sec), the immediate precursor of selenocysteinyl-tRNA(Sec). The sequence of Methanocaldococcus jannaschii PSTK (MjPSTK) suggests an N-terminal kinase domain (177 aa) followed by a presumed tRNA binding region (75 aa). The structures of MjPSTK complexed with ADP and AMPPNP revealed that this enzyme belongs to the P-loop kinase class, and that the kinase domain is closely related to gluconate kinase and adenylate kinase. ATP is bound by the P-loop domain (residues 11-18). Formed by antiparallel dimerization of two PSTK monomers, the enzyme structure shows a deep groove with positive electrostatic potential. Located in this groove is the enzyme's active site, which biochemical and genetic data suggest is composed of Asp-41, Arg-44, Glu-55, Tyr-82, Tyr-83, Met-86, and Met-132. Based on structural comparison with Escherichia coli adenylate kinase a docking model was generated that assigns these amino acids to the recognition of the terminal A76-Ser moieties of Ser-tRNA(Sec). The geometry and electrostatic environment of the groove in MjPSTK are perfectly complementary to the unusually long acceptor helix of tRNA(Sec).
Organizational Affiliation:
Department of Basic Medical Sciences, Institute of Medical Science, The University of Tokyo, 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639, Japan.