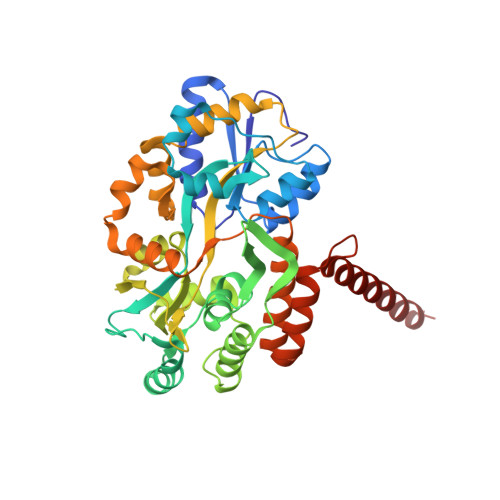
Mcm10 self-association is mediated by an N-terminal coiled-coil domain.
Du, W., Josephrajan, A., Adhikary, S., Bowles, T., Bielinsky, A.K., Eichman, B.F.(2013) PLoS One 8: e70518-e70518
- PubMed: 23894664
- DOI: https://doi.org/10.1371/journal.pone.0070518
- Primary Citation of Related Structures:
4JBZ - PubMed Abstract:
Minichromosome maintenance protein 10 (Mcm10) is an essential eukaryotic DNA-binding replication factor thought to serve as a scaffold to coordinate enzymatic activities within the replisome. Mcm10 appears to function as an oligomer rather than in its monomeric form (or rather than as a monomer). However, various orthologs have been found to contain 1, 2, 3, 4, or 6 subunits and thus, this issue has remained controversial. Here, we show that self-association of Xenopus laevis Mcm10 is mediated by a conserved coiled-coil (CC) motif within the N-terminal domain (NTD). Crystallographic analysis of the CC at 2.4 Å resolution revealed a three-helix bundle, consistent with the formation of both dimeric and trimeric Mcm10 CCs in solution. Mutation of the side chains at the subunit interface disrupted in vitro dimerization of both the CC and the NTD as monitored by analytical ultracentrifugation. In addition, the same mutations also impeded self-interaction of the full-length protein in vivo, as measured by yeast-two hybrid assays. We conclude that Mcm10 likely forms dimers or trimers to promote its diverse functions during DNA replication.
Organizational Affiliation:
Department of Biological Sciences, Vanderbilt University, Nashville, Tennessee, USA.