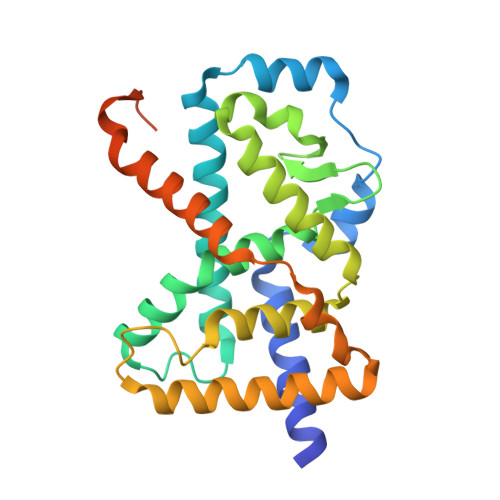
Ternary complex of human ROR gamma ligand-binding domain, inverse agonist and SMRT peptide shows a unique mechanism of corepressor recruitment
Noguchi, M., Nomura, A., Murase, K., Doi, S., Yamaguchi, K., Hirata, K., Shiozaki, M., Hirashima, S., Kotoku, M., Yamaguchi, T., Katsuda, Y., Steensma, R., Li, X., Tao, H., Tse, B., Fenn, M., Babine, R., Bradley, E., Crowe, P., Thacher, S., Adachi, T., Kamada, M.(2017) Genes Cells 22: 535-551
- PubMed: 28493531
- DOI: https://doi.org/10.1111/gtc.12494
- Primary Citation of Related Structures:
5X8Q, 5X8S, 5X8U, 5X8W, 5X8X - PubMed Abstract:
Retinoid-related orphan receptor gamma (RORγ) directly controls the differentiation of Th17 cell and the production of interleukin-17, which plays an integral role in autoimmune diseases. To obtain insight into RORγ, we have determined the first crystal structure of a ternary complex containing RORγ ligand-binding domain (LBD) bound with a novel synthetic inhibitor and a repressor peptide, 22-mer peptide from silencing mediator of retinoic acid and thyroid hormone receptor (SMRT). Comparison of a binary complex of nonliganded (apo) RORγ-LBD with a nuclear receptor co-activator (NCoA-1) peptide has shown that our inhibitor displays a unique mechanism different from those caused by natural inhibitor, ursolic acid (UA). The compound unprecedentedly induces indirect disruption of a hydrogen bond between His479 on helix 11 (H11) and Tyr502 on H12, which is crucial for active conformation. This crystallographic study will allow us to develop novel synthetic compounds for autoimmune disease therapy.
Organizational Affiliation:
Pharmaceutical Frontier Research Laboratories, Central Pharmaceutical Research Institute, Japan Tobacco Inc., 1-13-2, Fukuura, Kanazawa-Ku, Yokohama, Kanagawa, 236-0004, Japan.