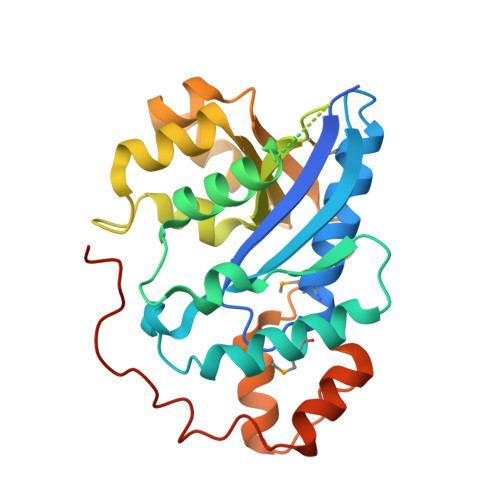
Crystal structure of human nicotinamide mononucleotide adenylyltransferase in complex with NMN.
Werner, E., Ziegler, M., Lerner, F., Schweiger, M., Heinemann, U.(2002) FEBS Lett 516: 239-244
- PubMed: 11959140
- DOI: https://doi.org/10.1016/s0014-5793(02)02556-5
- Primary Citation of Related Structures:
1GZU - PubMed Abstract:
The final step in the biosynthesis of nicotinamide-adenine dinucleotide, a major coenzyme in cellular redox reactions and involved in intracellular signaling, is catalyzed by the enzyme nicotinamide mononucleotide adenylyltransferase (NMNAT). The X-ray structure of human NMNAT in complex with nicotinamide mononucleotide was solved by the single-wavelength anomalous dispersion method at a resolution of 2.9 A. Human NMNAT is a symmetric hexamer whose subunit is formed by a large six-stranded parallel beta-sheet with helices on both sides. Human NMNAT displays a different oligomerization compared to the archaeal enzyme. The protein-nicotinamide mononucleotide interaction pattern provides insight into ligand binding in the human enzyme.
Organizational Affiliation:
Crystallography Group, Max Delbrück Center for Molecular Medicine, Robert-Rössle-Str. 10, D-13092, Berlin, Germany. ewe@mdc-berlin.de