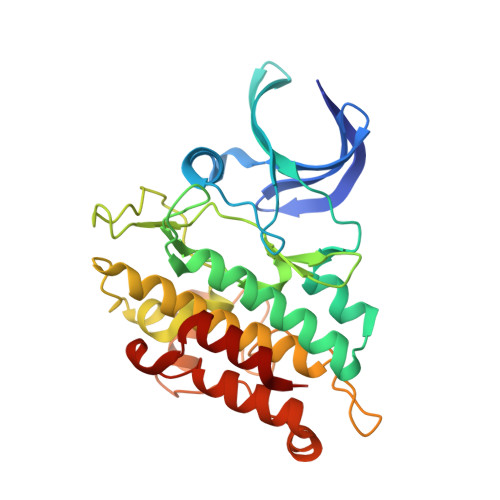
Crystal structure of the ACVR1 kinase domain in complex with the imidazo[1,2-b]pyridazine inhibitor K00135
Chaikuad, A., Sanvitale, C., Cooper, C., Canning, P., Mahajan, P., Daga, N., Petrie, K., Alfano, I., Gileadi, O., Fedorov, O., Krojer, T., Filippakopoulos, P., Muniz, J.R.C., von Delft, F., Weigelt, J., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Bullock, A., Structural Genomics Consortium (SGC)To be published.