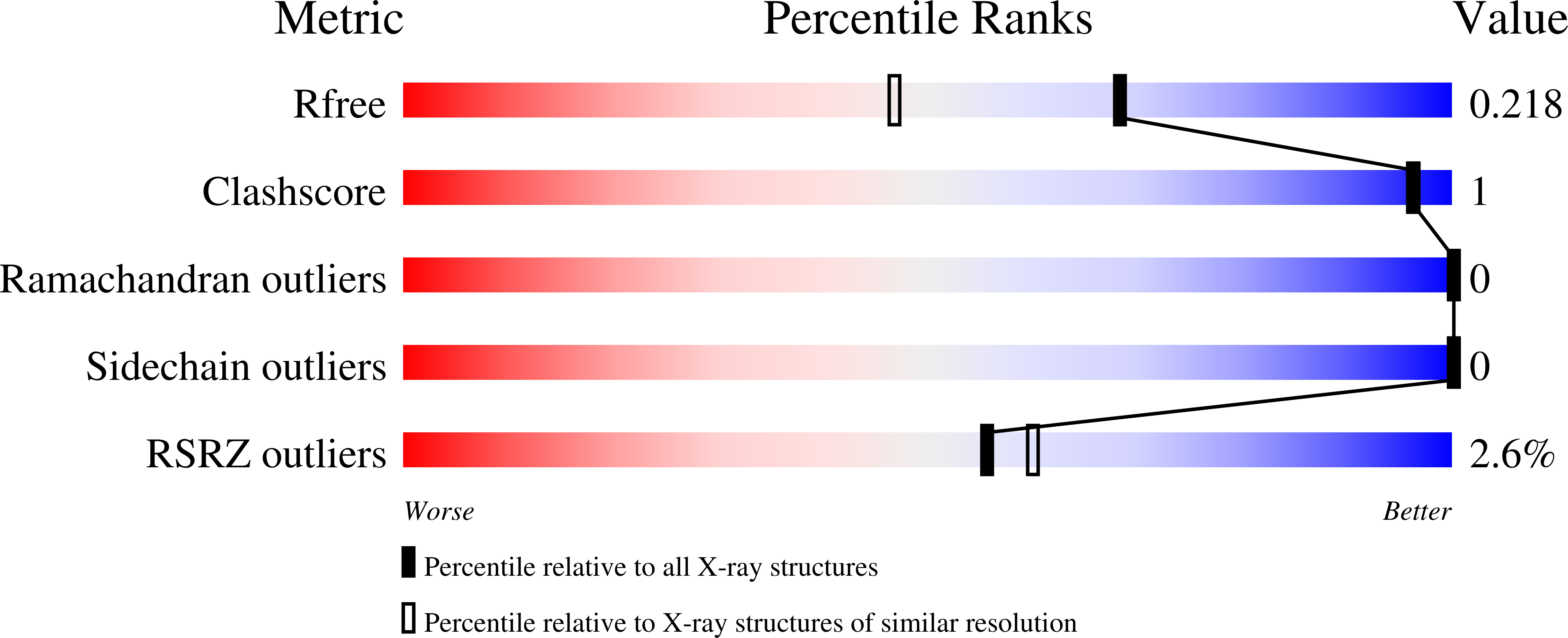
Pyrrolopyrimidine inhibitors of DNA gyrase B (GyrB) and topoisomerase IV (ParE). Part I: Structure guided discovery and optimization of dual targeting agents with potent, broad-spectrum enzymatic activity.
Tari, L.W., Trzoss, M., Bensen, D.C., Li, X., Chen, Z., Lam, T., Zhang, J., Creighton, C.J., Cunningham, M.L., Kwan, B., Stidham, M., Shaw, K.J., Lightstone, F.C., Wong, S.E., Nguyen, T.B., Nix, J., Finn, J.(2013) Bioorg Med Chem Lett 23: 1529-1536
- PubMed: 23352267
- DOI: https://doi.org/10.1016/j.bmcl.2012.11.032
- Primary Citation of Related Structures:
4GEE, 4GFN, 4GGL, 4HXW, 4HXZ, 4HY1, 4HYM, 4HYP, 4HZ0, 4HZ5 - PubMed Abstract:
The bacterial topoisomerases DNA gyrase (GyrB) and topoisomerase IV (ParE) are essential enzymes that control the topological state of DNA during replication. The high degree of conservation in the ATP-binding pockets of these enzymes make them appealing targets for broad-spectrum inhibitor development. A pyrrolopyrimidine scaffold was identified from a pharmacophore-based fragment screen with optimization potential. Structural characterization of inhibitor complexes conducted using selected GyrB/ParE orthologs aided in the identification of important steric, dynamic and compositional differences in the ATP-binding pockets of the targets, enabling the design of highly potent pyrrolopyrimidine inhibitors with broad enzymatic spectrum and dual targeting activity.
Organizational Affiliation:
Trius Therapeutics, 6310 Nancy Ridge Dr., San Diego, CA 92121, USA. ltari@triusrx.com