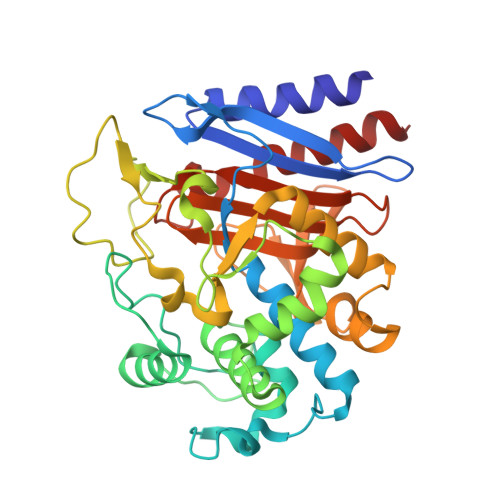
Structures of D-amino-acid amidase complexed with L-phenylalanine and with L-phenylalanine amide: insight into the D-stereospecificity of D-amino-acid amidase from Ochrobactrum anthropi SV3.
Okazaki, S., Suzuki, A., Mizushima, T., Komeda, H., Asano, Y., Yamane, T.(2008) Acta Crystallogr D Biol Crystallogr 64: 331-334
- PubMed: 18323628
- DOI: https://doi.org/10.1107/S0907444907067479
- Primary Citation of Related Structures:
2EFU, 2EFX - PubMed Abstract:
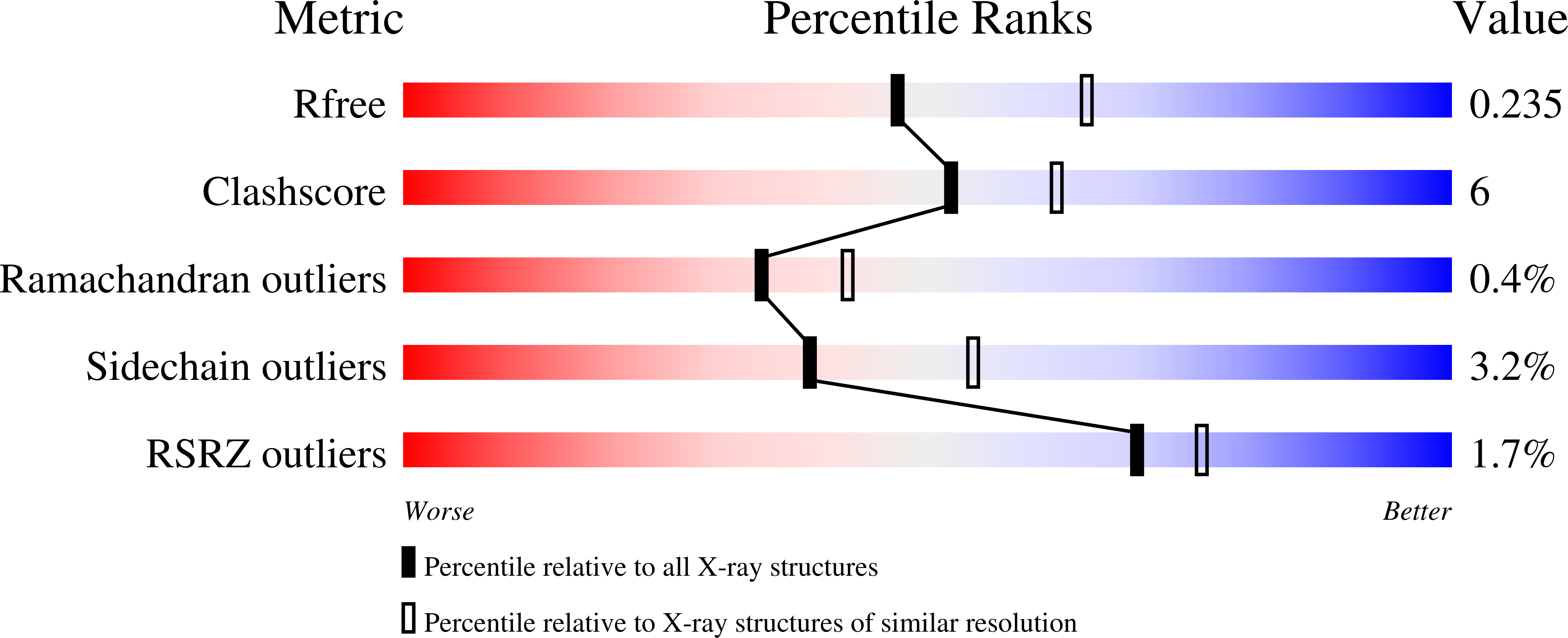
The crystal structures of D-amino-acid amidase (DAA) from Ochrobactrum anthropi SV3 in complex with L-phenylalanine and with L-phenylalanine amide were determined at 2.3 and 2.2 A resolution, respectively. Comparison of the L-phenylalanine amide complex with the D-phenylalanine complex reveals that the D-stereospecificity of DAA might be achieved as a consequence of three structural factors: (i) the hydrophobic cavity in the region in which the hydrophobic side chain of the substrate is held, (ii) the spatial arrangement of Gln310 O and Glu114 O epsilon2 that fixes the amino N atom of the substrate and (iii) the existence of two cavities that keep the carboxyl/amide group of the substrate near or apart from Ser60 O gamma.
Organizational Affiliation:
Department of Biotechnology, School of Engineering, Nagoya University, Chikusa, Nagoya 464-8603, Japan.