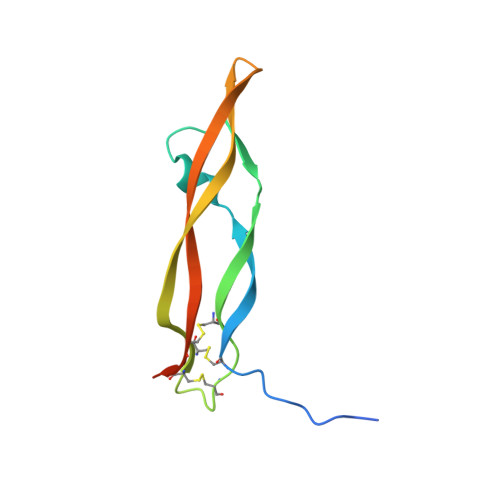
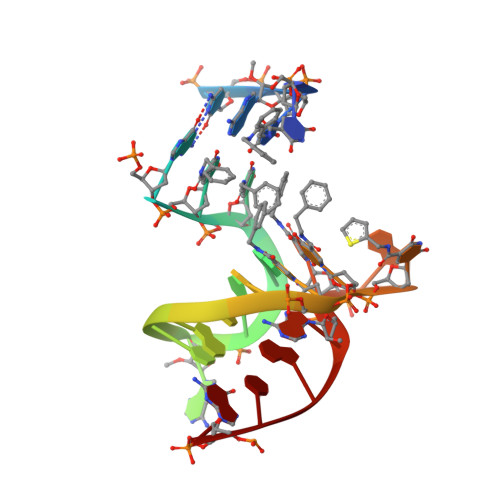
Unique motifs and hydrophobic interactions shape the binding of modified DNA ligands to protein targets.
Davies, D.R., Gelinas, A.D., Zhang, C., Rohloff, J.C., Carter, J.D., O'Connell, D., Waugh, S.M., Wolk, S.K., Mayfield, W.S., Burgin, A.B., Edwards, T.E., Stewart, L.J., Gold, L., Janjic, N., Jarvis, T.C.(2012) Proc Natl Acad Sci U S A 109: 19971-19976
- PubMed: 23139410
- DOI: https://doi.org/10.1073/pnas.1213933109
- Primary Citation of Related Structures:
4HQU, 4HQX - PubMed Abstract:
Selection of aptamers from nucleic acid libraries by in vitro evolution represents a powerful method of identifying high-affinity ligands for a broad range of molecular targets. Nevertheless, a sizeable fraction of proteins remain difficult targets due to inherently limited chemical diversity of nucleic acids. We have exploited synthetic nucleotide modifications that confer protein-like diversity on a nucleic acid scaffold, resulting in a new generation of binding reagents called SOMAmers (Slow Off-rate Modified Aptamers). Here we report a unique crystal structure of a SOMAmer bound to its target, platelet-derived growth factor B (PDGF-BB). The SOMAmer folds into a compact structure and exhibits a hydrophobic binding surface that mimics the interface between PDGF-BB and its receptor, contrasting sharply with mainly polar interactions seen in traditional protein-binding aptamers. The modified nucleotides circumvent the intrinsic diversity constraints of natural nucleic acids, thereby greatly expanding the structural vocabulary of nucleic acid ligands and considerably broadening the range of accessible protein targets.
Organizational Affiliation:
Emerald BioStructures, Inc, Bainbridge Island, WA 98110, USA.