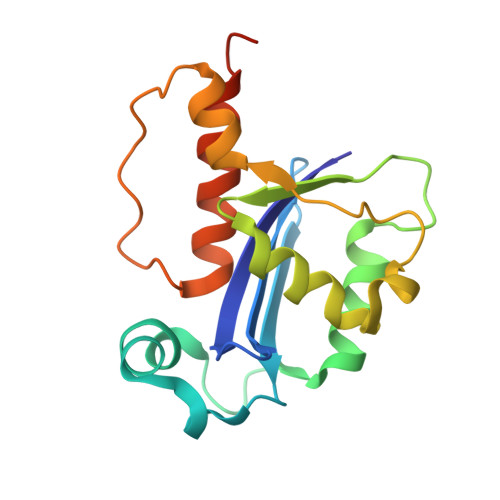
Structure and Function of RNase AS, a Polyadenylate-Specific Exoribonuclease Affecting Mycobacterial Virulence In Vivo
Romano, M., van de Weerd, R., Brouwer, F.C.C., Roviello, G.N., Lacroix, R., Sparrius, M., van den Brink-van Stempvoort, G., Maaskant, J.J., van der Sar, A.M., Appelmelk, B.J., Geurtsen, J.J., Berisio, R.(2014) Structure 22: 719-730
- PubMed: 24704253
- DOI: https://doi.org/10.1016/j.str.2014.01.014
- Primary Citation of Related Structures:
4OKE, 4OKJ, 4OKK - PubMed Abstract:
The cell-envelope of Mycobacterium tuberculosis plays a key role in bacterial virulence and antibiotic resistance. Little is known about the molecular mechanisms of regulation of cell-envelope formation. Here, we elucidate functional and structural properties of RNase AS, which modulates M. tuberculosis cell-envelope properties and strongly impacts bacterial virulence in vivo. The structure of RNase AS reveals a resemblance to RNase T from Escherichia coli, an RNase of the DEDD family involved in RNA maturation. We show that RNase AS acts as a 3'-5'-exoribonuclease that specifically hydrolyzes adenylate-containing RNA sequences. Also, crystal structures of complexes with AMP and UMP reveal the structural basis for the observed enzyme specificity. Notably, RNase AS shows a mechanism of substrate recruitment, based on the recognition of the hydrogen bond donor NH2 group of adenine. Our work opens a field for the design of drugs able to reduce bacterial virulence in vivo.
Organizational Affiliation:
Institute of Biostructures and Bioimaging, National Research Council, Via Mezzocannone 16, I-80134 Naples, Italy.