Welcome
RCSB Protein Data Bank (RCSB PDB) enables breakthroughs in science and education by providing access and tools for exploration, visualization, and analysis of:
Experimentally-determined 3D structures from the Protein Data Bank (PDB) archive | |
Computed Structure Models (CSM) from AlphaFold DB and ModelArchive |
These data can be explored in context of external annotations providing a structural view of biology.
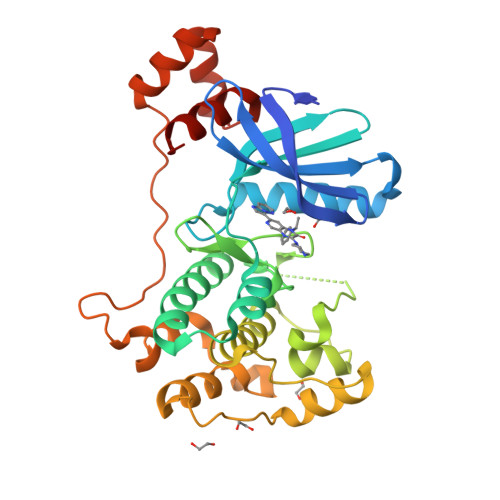
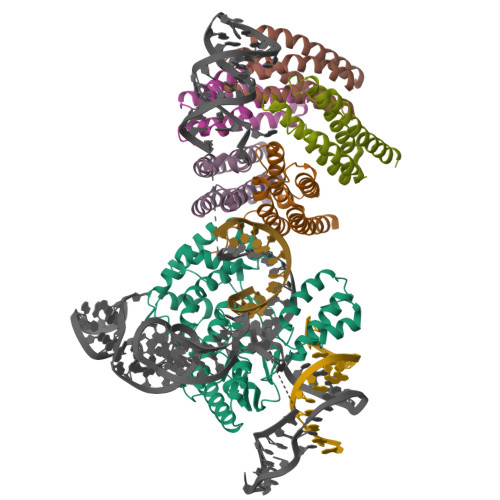
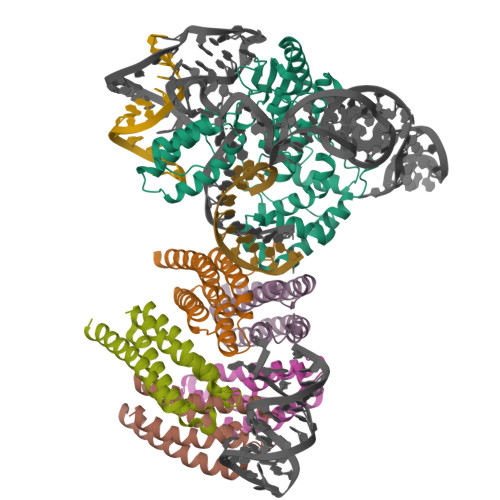
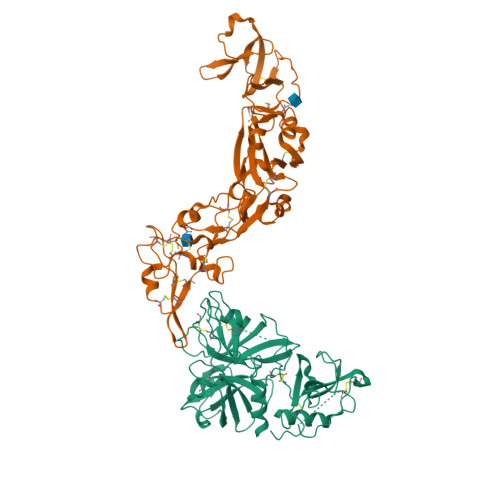
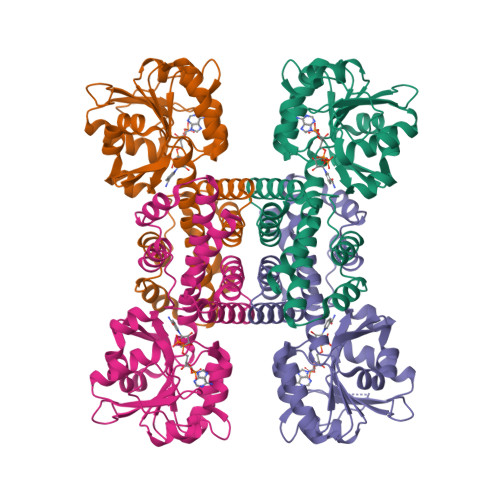
Latest Entries
Features & Highlights
Poster Prize Awarded at LACA
The wwPDB Foundation awarded Zaighum Abbas at the Latin American Crystallographic Association Meeting for The Role of SEPT7 in Magnaporthe oryzae: Structural Insights and Functional Implications
The Nobel Prize in Chemistry 2024
Congratulations to David Baker, who was recognized for computational protein design, and to Demis Hassabis and John M. Jumper, who were recognized for protein structure prediction. These achievements also celebrate the work of the PDB depositor community and the PDB archive that underpinned development of their prediction methods.
Register for the November 4 Mol* Virtual Office Hour
Join RCSB PDB for quick tips on how to use Mol* to view Sequence Annotations in 3D
wwPDB Data Release Policy on Public Preprint Archives
Contributions to public preprint archives that mention PDB/EMDB/BMRB entry IDs are deemed by the wwPDB to be publications
Register for the November 7 Virtual Office Hour on Supporting Extended PDB IDs
Interested in learning more about plans to extend PDB IDs to 12 characters? Bring you questions to a virtual Office Hour.
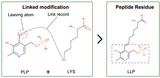
Annotation of Protein Modifications in the PDB
Protein chemical modifications (PCMs) and post translational modifications (PTMs) annotation is now included in CCD files, and updated atomic coordinate files are being rolled out.
Register Now for Webinar: Unlock Rapid Analyses Across the Whole PDB Using BinaryCIF
Join us on November 4 to future-proof your data analysis with BinaryCIF, a fully interchangeable yet drastically more efficient flavor of the PDBx/mmCIF format
Register for the October 8 Advanced Search Office Hour
Have questions about how to use RCSB.org? Join us for a virtual Office Hour.
Announcement: EDMAPS.rcsb.org Shutdown on October 16
Electron density map coefficients will instead be provided for all X-ray structures in the PDB archive.
Applications Open for Director
Distinguished scientists focused on 3D structures of biological macromolecules are invited to apply for this position with the RCSB PDB at Rutgers University.
See new feature archive
Interview with Director Emerita Helen Berman
Read about The huge protein database that spawned AlphaFold and biology's AI revolution in Nature» 10/20/2024Happy Birthday, Irving Geis
Celebrate Geis' birthday (October 18, 1908) with a tour of the Geis Digital Archive of his pioneering works of biomolecular art at PDB-101» 10/17/2024PDB-101 Focus: Peak Performance
PDB-101 materials explore the structural biology of athletics and well-being. Learn how the ten enzymes of glycolysis break down sugar in our diet in Glycolytic Enzymes» 10/15/2024Paper Published: MolViewSpec Toolkit
Learn about Describing and Sharing Molecular Visualizations Using the MolViewSpec Toolkit» 10/10/2024Explore Structural Biology with CSMs
Visit PDB-101 for more about Nobel Prize-winning Computed Structure Models and their measures of reliability; limitations; and how they can help determine experimental structures» 10/09/2024Fall Newsletter Published
Highlights Training Opportunities; Biocurator Milestone of >10,000 Depositions Processed; Influenza Virus painting; and more. In the Education Corner, learn about Paper Nucleic Acid Models for Hands-on Education» 10/07/2024Access IHM structures at wwPDB DOI Landing Pages
Structures determined by integrative and hybrid methods (IHM) are now available at wwPDB DOI landing pages alongside experimental structures in the PDB archive» 10/02/2024Explore Inktober SciArt images
Inktober is an annual ink drawing challenge to follow a list of drawing prompts and publish the work on social media. Structural biologist Irina Bezsonova (UCONN Health) created PDB-themed images for October in 2023 and 2022.» 09/30/2024Structural Biology and Nobel Prizes
Visit PDB-101 to explore connections between Nobel Prizes and more than 50 years of open access to the PDB archive.» 09/25/2024
Molecule of the Month

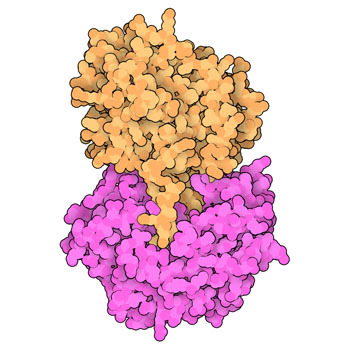
Angiotensin and Blood Pressure
Many medications for controlling high blood pressure inhibit the action of the peptide hormone angiotensin.
Read MoreQuarterly News (see archive)
Issue 103 - October 2024
In this Issue: Fall Training Opportunities; A Biocurator Milestone of >10,000 Depositions Processed; Influenza Virus painting; and more.
Education Corner: Paper Nucleic Acid Models for Hands-on Education by Cassandra K. Hayne and Phoebe A. Rice, The University of Chicago
Annual Reports

2023 Annual Report
Download the 2023 Annual Report (PDF) for an overview of recent activities, including RCSB PDB APIs and pairwise alignment tools.