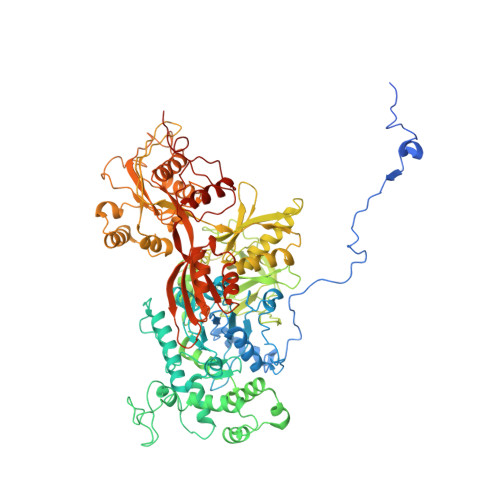
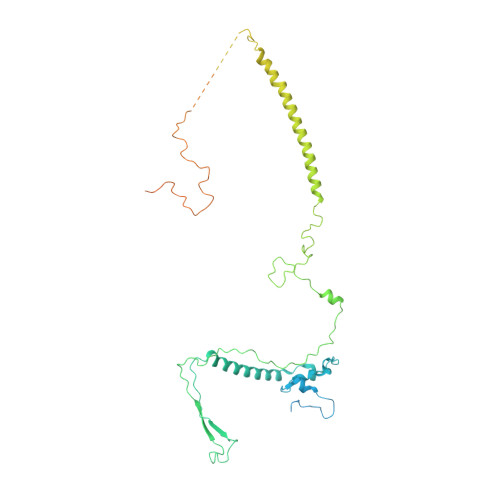
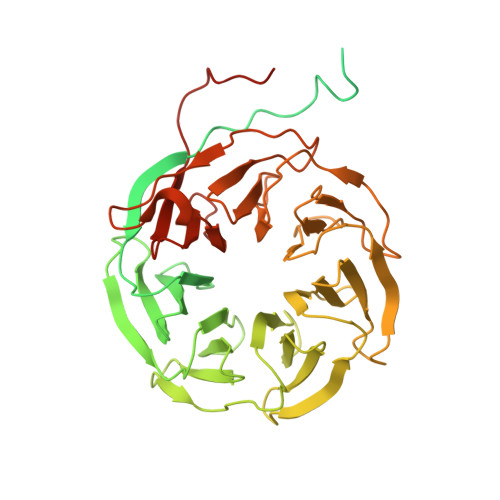
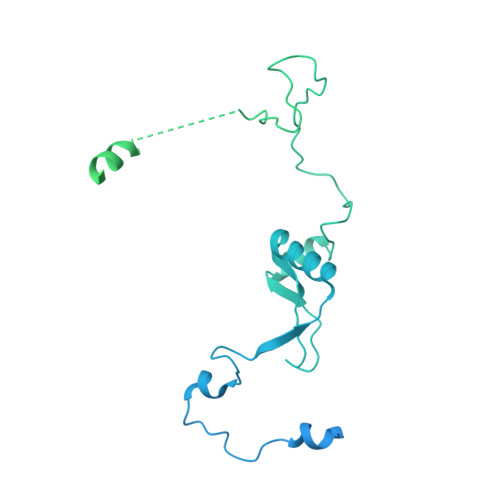
Structure and Conformational Dynamics of the Human Spliceosomal BactComplex.
Haselbach, D., Komarov, I., Agafonov, D.E., Hartmuth, K., Graf, B., Dybkov, O., Urlaub, H., Kastner, B., Luhrmann, R., Stark, H.(2018) Cell 172: 454-464.e11
- PubMed: 29361316
- DOI: https://doi.org/10.1016/j.cell.2018.01.010
- Primary Citation of Related Structures:
6FF4, 6FF7 - PubMed Abstract:
The spliceosome is a highly dynamic macromolecular complex that precisely excises introns from pre-mRNA. Here we report the cryo-EM 3D structure of the human B act spliceosome at 3.4 Å resolution. In the B act state, the spliceosome is activated but not catalytically primed, so that it is functionally blocked prior to the first catalytic step of splicing. The spliceosomal core is similar to the yeast B act spliceosome; important differences include the presence of the RNA helicase aquarius and peptidyl prolyl isomerases. To examine the overall dynamic behavior of the purified spliceosome, we developed a principal component analysis-based approach. Calculating the energy landscape revealed eight major conformational states, which we refined to higher resolution. Conformational differences of the highly flexible structural components between these eight states reveal how spliceosomal components contribute to the assembly of the spliceosome, allowing it to generate a dynamic interaction network required for its subsequent catalytic activation.
- Department for Structural Dynamics, Max Planck Institute for Biophysical Chemistry, Am Fassberg 11, 37077 Göttingen, Germany.
Organizational Affiliation: