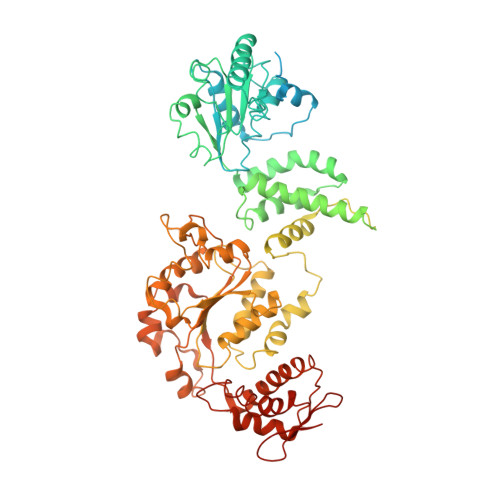
Structural basis for substrate gripping and translocation by the ClpB AAA+ disaggregase.
Rizo, A.N., Lin, J., Gates, S.N., Tse, E., Bart, S.M., Castellano, L.M., DiMaio, F., Shorter, J., Southworth, D.R.(2019) Nat Commun 10: 2393-2393
- PubMed: 31160557
- DOI: https://doi.org/10.1038/s41467-019-10150-y
- Primary Citation of Related Structures:
6OAX, 6OAY, 6OG1, 6OG2, 6OG3 - PubMed Abstract:
Bacterial ClpB and yeast Hsp104 are homologous Hsp100 protein disaggregases that serve critical functions in proteostasis by solubilizing protein aggregates. Two AAA+ nucleotide binding domains (NBDs) power polypeptide translocation through a central channel comprised of a hexameric spiral of protomers that contact substrate via conserved pore-loop interactions. Here we report cryo-EM structures of a hyperactive ClpB variant bound to the model substrate, casein in the presence of slowly hydrolysable ATPγS, which reveal the translocation mechanism. Distinct substrate-gripping interactions are identified for NBD1 and NBD2 pore loops. A trimer of N-terminal domains define a channel entrance that binds the polypeptide substrate adjacent to the topmost NBD1 contact. NBD conformations at the seam interface reveal how ATP hydrolysis-driven substrate disengagement and re-binding are precisely tuned to drive a directional, stepwise translocation cycle.
Organizational Affiliation:
Graduate Program in Chemical Biology, University of Michigan, Ann Arbor, MI, 48109, USA.