Leishmania major N-myristoyltransferase in complex with indazole inhibitor IMP-918
Brannigan, J.A.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glycylpeptide N-tetradecanoyltransferase | A [auth AAA] | 418 | Leishmania major | Mutation(s): 0 Gene Names: NMT, LMJF_32_0080 EC: 2.3.1.97 | 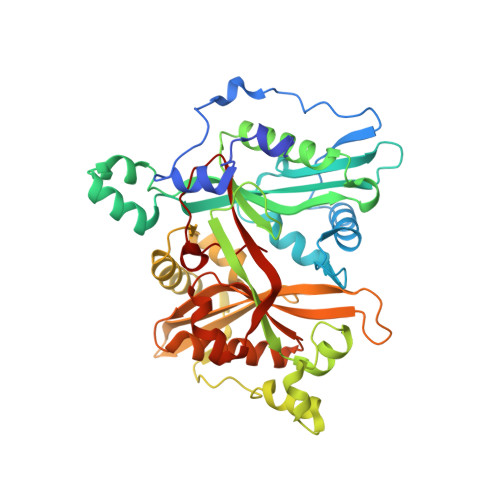 |
UniProt | |||||
Find proteins for Q4Q5S8 (Leishmania major) Explore Q4Q5S8 Go to UniProtKB: Q4Q5S8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q4Q5S8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MYA Query on MYA | B [auth AAA] | TETRADECANOYL-COA C35 H62 N7 O17 P3 S DUAFKXOFBZQTQE-QSGBVPJFSA-N |  | ||
| NZB (Subject of Investigation/LOI) Query on NZB | D [auth AAA] | [(5-{4-fluoro-2-[2-(pyridin-3-yl)ethoxy]phenyl}-1H-indazol-3-yl)methyl]dimethylamine C23 H23 F N4 O RNWBRNZKOWMEPZ-UHFFFAOYSA-N |  | ||
| MG Query on MG | C [auth AAA] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 48.379 | α = 90 |
| b = 91.899 | β = 113.697 |
| c = 53.609 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| SCALA | data scaling |
| REFMAC | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Wellcome Trust | -- |