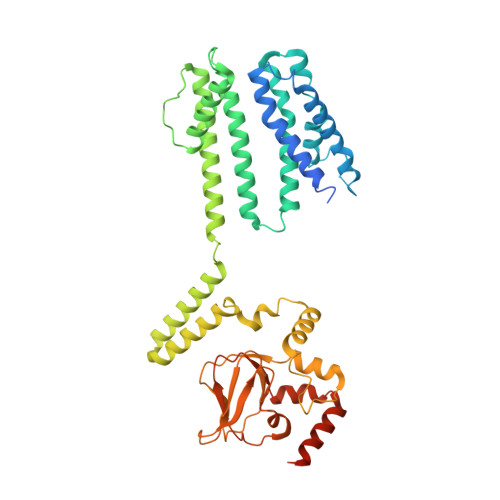
Gating intermediates reveal inhibitory role of the voltage sensor in a cyclic nucleotide-modulated ion channel.
Gao, X., Schmidpeter, P.A.M., Berka, V., Durham, R.J., Fan, C., Jayaraman, V., Nimigean, C.M.(2022) Nat Commun 13: 6919-6919
- PubMed: 36376326
- DOI: https://doi.org/10.1038/s41467-022-34673-z
- Primary Citation of Related Structures:
7RSH, 7RTF, 7RTJ, 7RU0, 7RYR, 7RYS - PubMed Abstract:
Understanding how ion channels gate is important for elucidating their physiological roles and targeting them in pathophysiological states. Here, we used SthK, a cyclic nucleotide-modulated channel from Spirochaeta thermophila, to define a ligand-gating trajectory that includes multiple on-pathway intermediates. cAMP is a poor partial agonist for SthK and depolarization increases SthK activity. Tuning the energy landscape by gain-of-function mutations in the voltage sensor domain (VSD) allowed us to capture multiple intermediates along the ligand-activation pathway, highlighting the allosteric linkage between VSD, cyclic nucleotide-binding (CNBD) and pore domains. Small, lateral displacements of the VSD S4 segment were necessary to open the intracellular gate, pointing to an inhibitory VSD at rest. We propose that in wild-type SthK, depolarization leads to such VSD displacements resulting in release of inhibition. In summary, we report conformational transitions along the activation pathway that reveal allosteric couplings between key sites integrating to open the intracellular gate.
- Department of Anesthesiology, Weill Cornell Medical College, 1300 York Avenue, New York, NY, 10065, USA.
Organizational Affiliation: