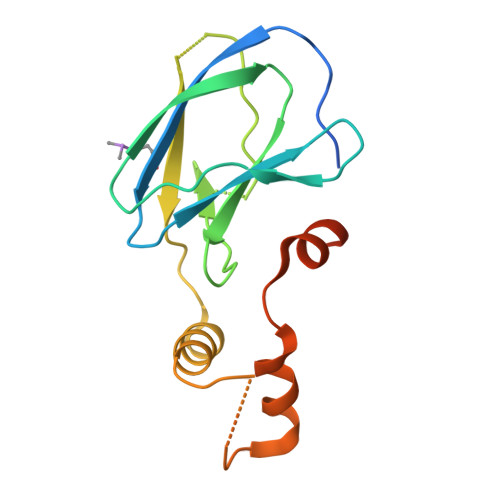
Identification of novel 7-hydroxycoumarin derivatives as ELOC binders with potential to modulate CRL2 complex formation.
Kim, Y., Baek, S.J., Yoon, E.K., Choi, M., Kim, J.H., Kim, K., Park, C.H., Lee, B.I.(2025) Sci Rep 15: 3622-3622
- PubMed: 39881207
- DOI: https://doi.org/10.1038/s41598-025-88166-2
- Primary Citation of Related Structures:
8ZV8, 8ZVJ, 9IPW - PubMed Abstract:
The VHL-containing cullin-RING E3 ubiquitin ligase (CRL2 VHL ) complex is an E3 ligase commonly used in targeted protein degradation (TPD). Hydroxyproline-based ligands that mimic VHL substrates have been developed as anchor molecules for proteolysis-targeting chimeras (PROTACs) in TPD. To expand the chemical space for VHL ligands, we conducted fragment screening using VHL-ELOB-ELOC (VBC) proteins. We found that certain 7-hydroxycoumarin derivatives (7HCs), rather than VHL, would bind to the ELOC component of the VBC complex. The 7HC binding site overlapped with the CUL2 binding interface on ELOC but did not overlap with the CUL5 binding interface, suggesting that 7HCs may influence the formation of CRL2 but not CRL5. Although the binding affinities of these 7HCs to the VBC complex were relatively low, they represent novel and promising foundational agents for the development of chemical probes or inhibitors that target ELOC-containing CRLs.
- Research Institute, National Cancer Center, Goyang-si, 10408, Gyeonggi, Republic of Korea.
Organizational Affiliation: