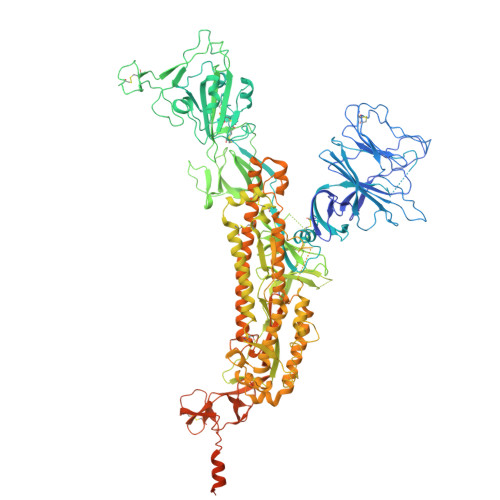
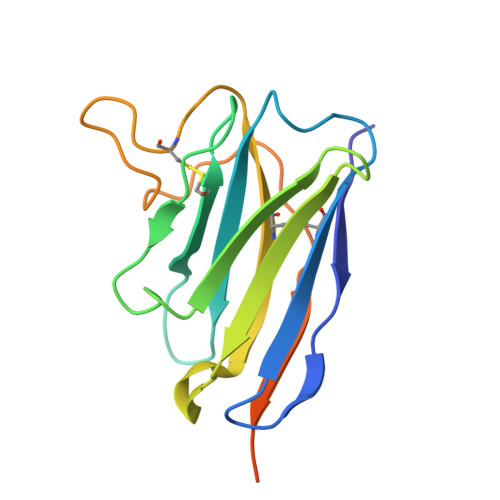
Discovery of Nanosota-9 as anti-Omicron nanobody therapeutic candidate.
Ye, G., Bu, F., Saxena, D., Turner-Hubbard, H., Herbst, M., Spiller, B., Wadzinski, B.E., Du, L., Liu, B., Zheng, J., Li, F.(2024) PLoS Pathog 20: e1012726-e1012726
- PubMed: 39591462
- DOI: https://doi.org/10.1371/journal.ppat.1012726
- Primary Citation of Related Structures:
9CO6, 9CO7, 9CO8, 9CO9 - PubMed Abstract:
Omicron subvariants of SARS-CoV-2 continue to pose a significant global health threat. Nanobodies, single-domain antibodies derived from camelids, are promising therapeutic tools against pandemic viruses due to their favorable properties. In this study, we identified a novel nanobody, Nanosota-9, which demonstrates high potency against a wide range of Omicron subvariants both in vitro and in a mouse model. Cryo-EM data revealed that Nanosota-9 neutralizes Omicron through a unique mechanism: two Nanosota-9 molecules crosslink two receptor-binding domains (RBDs) of the trimeric Omicron spike protein, preventing the RBDs from binding to the ACE2 receptor. This mechanism explains its strong anti-Omicron potency. Additionally, the Nanosota-9 binding epitopes on the spike protein are relatively conserved among Omicron subvariants, contributing to its broad anti-Omicron spectrum. Combined with our recently developed structure-guided in vitro evolution approach for nanobodies, Nanosota-9 has the potential to serve as the foundation for a superior anti-Omicron therapeutic.
- Department of Pharmacology, University of Minnesota Medical School, Minneapolis, Minnesota, United States of America.
Organizational Affiliation: