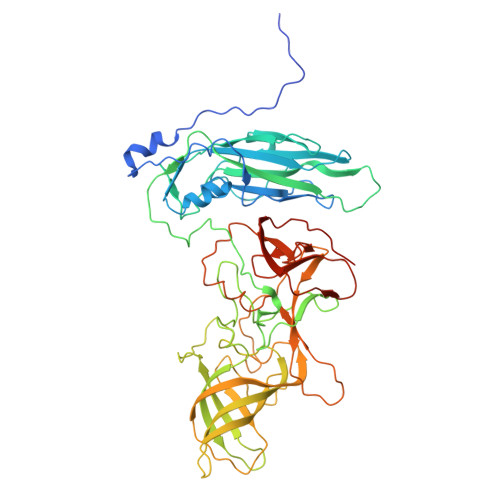
Murine norovirus allosteric escape mutants mimic gut activation.
Sherman, M.B., Smith, H.Q., Cox, F., Wobus, C.E., Lynch, G.C., Pettitt, B.M., Smith, T.J.(2025) J Virol 99: e0021925-e0021925
- PubMed: 40353669
- DOI: https://doi.org/10.1128/jvi.00219-25
- Primary Citation of Related Structures:
9EAN, 9EAO, 9EAP, 9EAQ - PubMed Abstract:
Murine norovirus (MNV) undergoes large conformational changes in response to the environment. The T=3 icosahedral capsid is composed of 180 copies of ~58 kDa VP1 that has N-terminal (N), shell (S), and C-terminal protruding (P) domains. In phosphate-buffered saline, the P domains are loosely tethered to the shell and float ~15 Å above the surface. At conditions found in the gut (i.e., low pH with high metal ion and bile salt concentrations), the P domain rotates and drops onto the shell with intra P domain changes that enhance receptor interactions while blocking antibody binding. Two of our monoclonal antibodies (2D3 and 4F9) have broad strain recognition, and the only escape mutants, V339I and D348E, are located on the C'D' loop and ~20 Å from the epitope. Here, we determined the cryo-EM structures of V339I and D348E at neutral pH +/-metal ions and bile salts. These allosteric escape mutants have the activated conformation in the absence of gut triggers. Since this conformation is not recognized by antibodies, it explains how these mutants evade antibody recognition. Dynamic simulations of the P domain further suggest that movement of the C'D' loop may be the rate-limiting step in the conformational change and that V339I increases the motion of the A'B'/E'F' loops compared to the wild-type (WT), facilitating the transition to the activated state. These findings have important implications for norovirus vaccine design since they uncover a form of the viral capsid that should lend superior immune protection against subsequent challenge by wild-type virus.IMPORTANCEImmune protection from norovirus infection is notoriously transient in both humans and mice. Our results strongly suggest that this is likely because the "activated" form of the virus found in gut conditions is not recognized by antibodies created in the circulation. By reversibly presenting one structure in the gut and a completely different antigenic structure in circulation, the gut tissue can be infected in subsequent challenges, while extraintestinal organs are protected. We find here that allosteric escape mutants to the most broadly neutralizing antibodies thwart recognition by transitioning to the activated state without the need for gut triggers (i.e., bile, low pH, or metal ions). These findings are significant because it is now feasible to present the activated form of the virus to the immune system (for example, as a vaccine) to better protect the gut tissue for longer periods of time.
- Department of Biochemistry and Molecular Biology, University of Texas Medical Branch at Galveston, Galveston, Texas, USA.
Organizational Affiliation: