Structural insights into mechanism of activation and deactivation of prostaglandin receptors
Davoudinasab, B., Han, G.W., Cherezov, V.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| anti-BRIL Fab Heavy Chain | A [auth H] | 229 | Homo sapiens | Mutation(s): 0 |  |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| anti-Fab Nanobody synthetic construct | B [auth K] | 123 | synthetic construct | Mutation(s): 0 |  |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| anti-BRIL Fab Light Chain | C [auth L] | 214 | synthetic construct | Mutation(s): 0 | 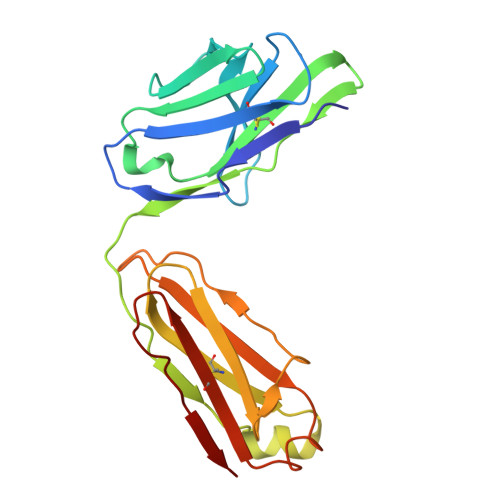 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Prostaglandin D2 receptor,Soluble cytochrome b562 | D [auth A] | 495 | Homo sapiens | Mutation(s): 6 Gene Names: PTGDR, cybC | 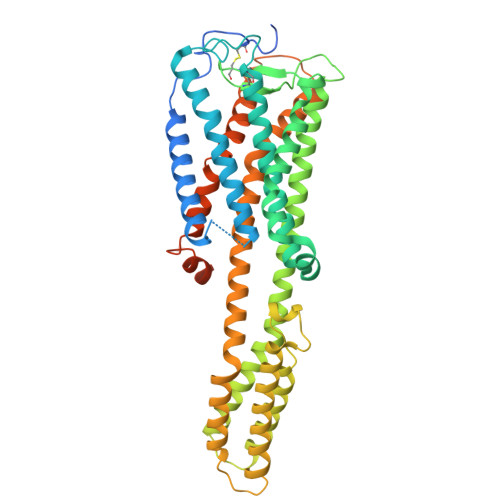 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P0ABE7 (Escherichia coli) Explore P0ABE7 Go to UniProtKB: P0ABE7 | |||||
Find proteins for Q13258 (Homo sapiens) Explore Q13258 Go to UniProtKB: Q13258 | |||||
PHAROS: Q13258 GTEx: ENSG00000168229 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | Q13258P0ABE7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1BIL (Subject of Investigation/LOI) Query on A1BIL | E [auth A] | [2-chloro-5-(2,6-dimethyl-4-{[(2S)-4-methyl-3,4-dihydro-2H-1,4-benzoxazin-2-yl]methoxy}benzamido)phenyl]acetic acid C27 H27 Cl N2 O5 KHJQYLXBPSJLGG-NRFANRHFSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| RECONSTRUCTION | cryoSPARC | 4.7.0 |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | R35GM127086 |