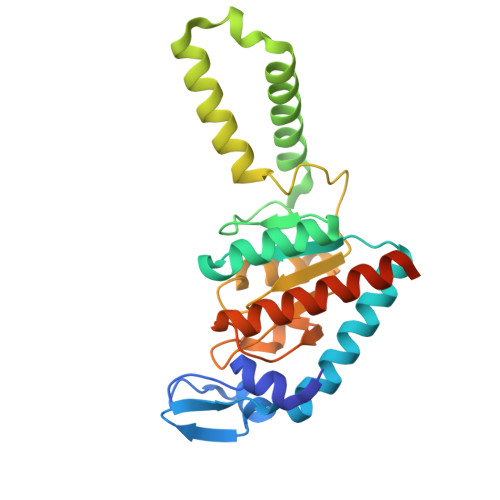
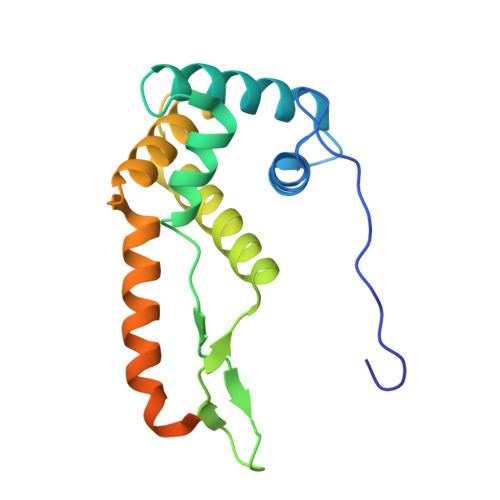
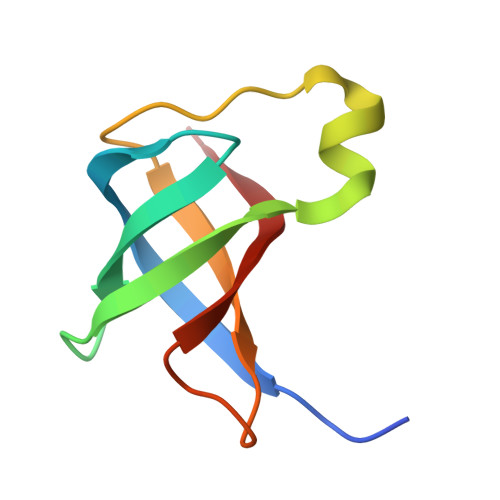
A binding site for the antibiotic GE81112 in the ribosomal mRNA channel.
Schedlbauer, A., Han, X., van Bakel, W., Kaminishi, T., Ochoa-Lizarralde, B., Iturrioz, I., Capuni, R., Parry, R., Zegarra, R., Gil-Carton, D., Lopez-Alonso, J.P., Barragan Sanz, K., Brandi, L., Gualerzi, C.O., Fucini, P., Connell, S.R.(2024) bioRxiv
- PubMed: 39386670
- DOI: https://doi.org/10.1101/2024.09.26.614503
- Primary Citation of Related Structures:
9H8G, 9H9H, 9H9I, 9H9J, 9H9K, 9H9L, 9H9M, 9H9N - PubMed Abstract:
The initiation phase is the rate-limiting step of protein synthesis (translation) and is finely regulated, making it an important drug target. In bacteria, initiation is guided by three initiation factors and involves positioning the start site on the messenger RNA within the P-site on the small ribosomal subunit (30S), where it is decoded by the initiator tRNA. This process can be efficiently inhibited by GE81112, a natural hydrophilic, noncyclic, nonribosomal tetrapeptide. It is found in nature in three structural variants (A, B and B1 with molecular masses of 643-658 Da). Previous biochemical and structural characterisation of GE81112 indicates that the primary mechanism of action of this antibiotic is to (1) prevent the initiator tRNA from binding correctly to the P-site and (2) block conformational rearrangements in initiation factor IF3, resulting in an unlocked 30S pre/C state. In this study, using cryoEM, we have determined the binding site of GE81112 in initiation complexes (3.2-3.7Å) and on empty ribosomes (2.09 Å). This binding site is within the mRNA channel (E-site) but remote from the binding site of the initiation factors and initiator tRNA. This suggests that it acts allosterically to prevent the initiator tRNA from being locked into place. The binding mode is consistent with previous biochemical studies and recent work identifying the key pharmacophores of GE81112.
- Center for Cooperative Research in Biosciences (CIC bioGUNE), Basque Research and Technology Alliance (BRTA), Bizkaia Technology Park, Building 801A, 48160 Derio, Spain.
Organizational Affiliation: