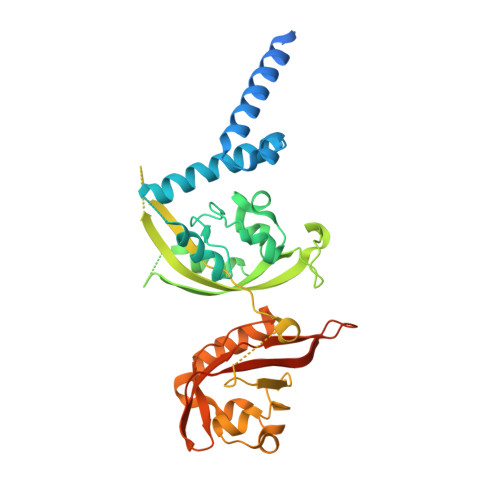
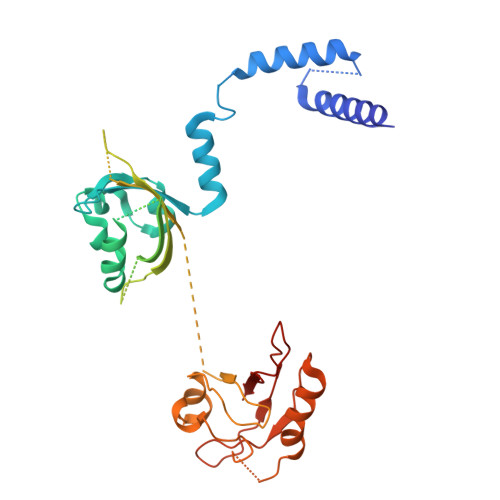
Context-Dependent Variability Of HIF Heterodimers Influences Interactions With Macromolecular And Small Molecule Partners.
Closson, J.D., Xu, X., Zhang, M., Tiyani, T.T., Marcelino, L.P., Isiorho, E.A., Nagati, J.S., Garcia, J.A., Gardner, K.H.(2025) bioRxiv
- PubMed: 40502054
- DOI: https://doi.org/10.1101/2025.05.29.656908
- Primary Citation of Related Structures:
9OF0, 9OF2, 9OFU, 9OPF - PubMed Abstract:
Hypoxia inducible factors (HIFs) are transcription factors that coordinate cellular responses to low oxygen levels, functioning as an α/β heterodimer which binds a short hypoxia response element (HRE) DNA sequence. Prior studies suggest HIF/HRE complexes are augmented by the binding of additional factors nearby, but those interactions are not well understood. Here, we integrated structural and biochemical approaches to investigate several functionally relevant HIF assemblies with other protein, small molecule, and DNA partners. First, we used cryo-electron microscopy (cryo-EM) to establish HIF-1 and HIF-2 form novel "dimer-of-heterodimers" (DoHD) complexes on extended human EPO enhancer sequences, showing that one heterodimer bound at a canonical HRE site with the second binding in an inverted fashion to an HRE-adjacent sequence (HAS) 8 bp away. Consistent with ARNT PAS-B domains predominating interactions within a DoHD, we found HIF-1 and HIF-2 assemble mixed DoHD complexes on the same DNA. Second, we saw substantial variability among ligands for isolated ARNT or HIF-2α PAS-B domains to bind larger complexes: for example, the ARNT PAS-B binding KG-548 and KG-279 ligands both bound the simpler HIF-2 heterodimer but exhibited differential binding to a HIF-2 DoHD. Finally, we combined cryo-EM and hydrogen-deuterium exchange by mass spectrometry (HDX-MS) to show how HIF-1 and HIF-2 heterodimers engage the transforming acidic coiled-coil containing protein 3 (TACC3) coactivator via both ARNT and HIF-α subunits, though this was unseen in the larger DoHD. Our findings highlight the importance of both molecular context and dynamics in biomolecular complex formation, adding to the complexities of potential regulation. Hypoxia inducible factors (HIFs) are transcription factors that regulate oxygen-dependent cellular processes with implications in certain types of cancers. Current molecular structures of HIFs bound to short DNA fragments provide insights into their function, but leave open questions about how they bind longer natural DNA fragments and interact with small molecules and protein coactivators. Integrating structural and biochemical techniques, we discovered a novel assembly in which two HIFs bind together on a single extended DNA fragment, forming a "dimer-of-heterodimers", and ascertained how structural differences arising from higher-ordered complex formation affect ligand and coactivator binding. Our studies highlight how functional contexts can shift structural paradigms and provide greater insight into the mechanisms by which HIFs and similar transcription factors operate.