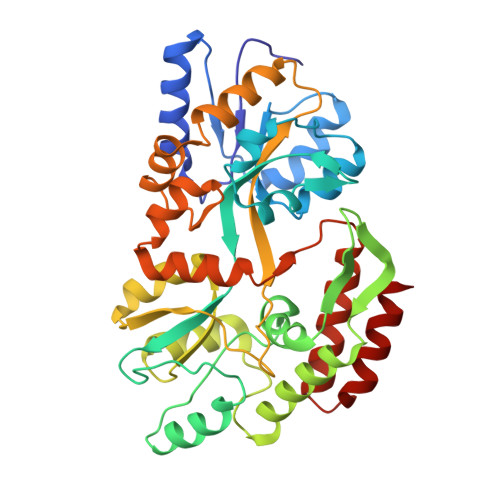
Refined 1.8-A structure reveals the mode of binding of beta-cyclodextrin to the maltodextrin binding protein.
Sharff, A.J., Rodseth, L.E., Quiocho, F.A.(1993) Biochemistry 32: 10553-10559
- PubMed: 8399200
- DOI: https://doi.org/10.1021/bi00091a004
- Primary Citation of Related Structures:
1DMB - PubMed Abstract:
The maltodextrin binding protein from Escherichia coli serves as the initial receptor for both the active transport of and chemotaxis toward a range of linear maltose sugars. The X-ray structures of both the maltose-bound and sugar-free forms of the protein have been previously described [Spurlino, J. C., Lu, G.-Y., & Quiocho, F. A. (1991) J. Biol. Chem. 266, 5202-5219; Sharff, A. J., Rodseth, L. E., Spurlino, J. C., & Quocho, F. A. (1992) Biochemistry 31, 10657-10663]. The X-ray crystal structure of the maltodextrin binding protein complexed with cyclomaltoheptaose (beta-cyclodextrin) has been determined from a single crystal. The structure has been refined to a final R-value of 21% at 1.8-A resolution. Although not a physiological ligand for the maltodextrin binding protein, beta-cyclodextrin has been shown to bind with a Kd of the same order as those of the linear maltodextrin substrates. The observed structure shows that the complexed protein remains in the fully open conformation and is almost identical to the structure of the unliganded protein. The sugar sits in the open cleft with three glucosyl units bound to the C-domain at the base of the cleft, in a similar position to maltotriose, the most tightly bound ligand. The top of the ring is loosely bound to the upper edge of the cleft on the N-domain. The sugar makes a total of 94 productive interactions (of less than 4.0-A length) with the protein and with bound water molecules.(ABSTRACT TRUNCATED AT 250 WORDS)
Organizational Affiliation:
Howard Hughes Medical Institute, Baylor College of Medicine, Houston, Texas 77030.