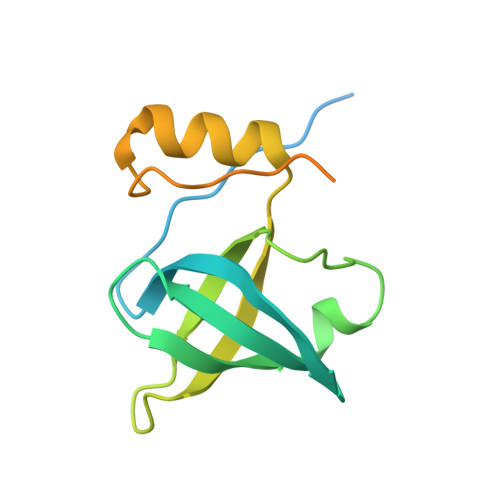
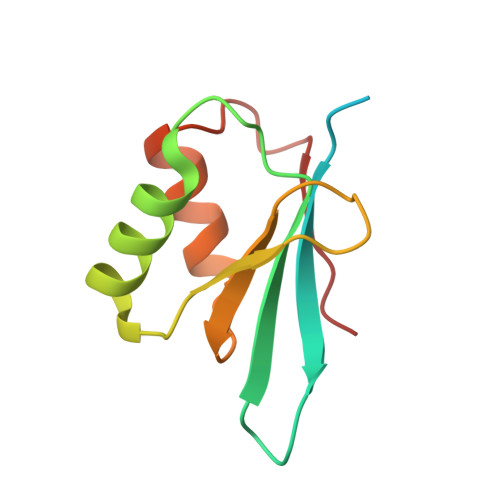
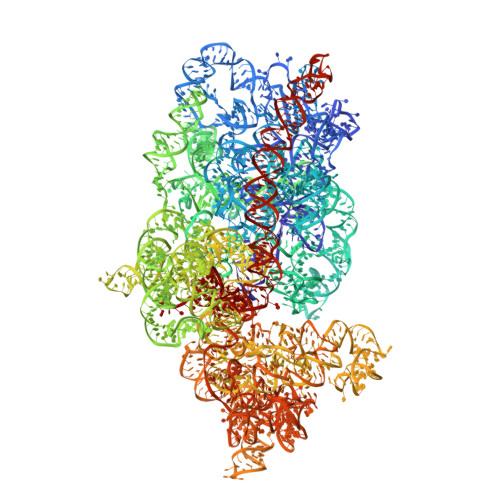
Conformational Differences between Open and Closed States of the Eukaryotic Translation Initiation Complex.
Llacer, J.L., Hussain, T., Marler, L., Aitken, C.E., Thakur, A., Lorsch, J.R., Hinnebusch, A.G., Ramakrishnan, V.(2015) Mol Cell 59: 399-412
- PubMed: 26212456
- DOI: https://doi.org/10.1016/j.molcel.2015.06.033
- Primary Citation of Related Structures:
3JAM, 3JAP - PubMed Abstract:
Translation initiation in eukaryotes begins with the formation of a pre-initiation complex (PIC) containing the 40S ribosomal subunit, eIF1, eIF1A, eIF3, ternary complex (eIF2-GTP-Met-tRNAi), and eIF5. The PIC, in an open conformation, attaches to the 5' end of the mRNA and scans to locate the start codon, whereupon it closes to arrest scanning. We present single particle cryo-electron microscopy (cryo-EM) reconstructions of 48S PICs from yeast in these open and closed states, at 6.0 Å and 4.9 Å, respectively. These reconstructions show eIF2β as well as a configuration of eIF3 that appears to encircle the 40S, occupying part of the subunit interface. Comparison of the complexes reveals a large conformational change in the 40S head from an open mRNA latch conformation to a closed one that constricts the mRNA entry channel and narrows the P site to enclose tRNAi, thus elucidating key events in start codon recognition.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Cambridge CB2 0QH, UK.