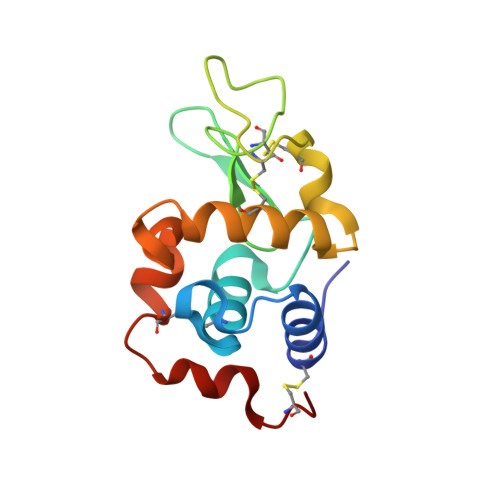
An ensemble of crystallographic models enables the description of novel bromate-oxoanion species trapped within a protein crystal
Ondracek, J., Mesters, J.R.(2006) Acta Crystallogr D Biol Crystallogr 62: 996-1001
- PubMed: 16929100
- DOI: https://doi.org/10.1107/S0907444906021627
- Primary Citation of Related Structures:
2D6B - PubMed Abstract:
Only a few protein-oxoanion crystal complexes have been described to date. Here, the structure of a protein soaked in a bromate solution has been determined to a resolution of 1.25 A and refined to final overall R/R(free) values of 18.04/21.3 (isotropic) and 11.25/14.67 (anisotropic). In contrast to the single-model approach, refinement of an ensemble of ten models enabled us to determine variances and statistically evaluate bond-length distances and angles in the oxoanions. In total, nine bromate positions, including two BrO(3)(-) x HBrO(3) dimer species, have been identified on the basis of the anomalous signal of the Br atoms. For all bromate ions, the main-chain amide atoms of the protein were identified as the dominant binding positions, a useful property in any experimental phase-determination experiment.
- Department of Recombinant Expression and Structural Biology, Institute of Molecular Genetics, Academy of Sciences of the Czech Republic, Flemingovo namesti 2, CZ-16637 Praha 6, Czech Republic. ondracek@img.cas.cz
Organizational Affiliation: