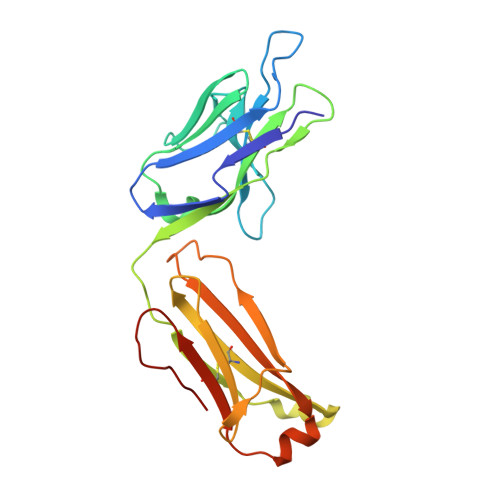
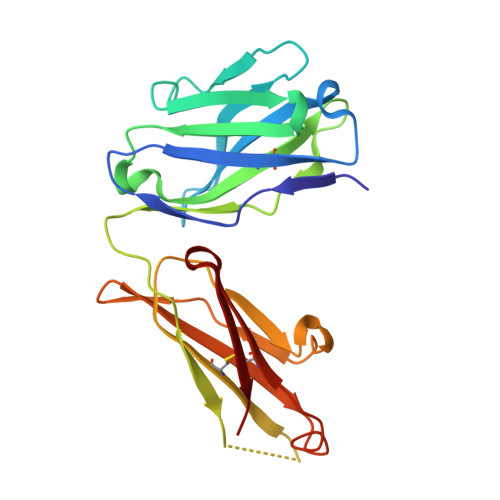
Structure of anti-FLAG M2 Fab domain and its use in the stabilization of engineered membrane proteins.
Roosild, T.P., Castronovo, S., Choe, S.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 835-839
- PubMed: 16946459
- DOI: https://doi.org/10.1107/S1744309106029125
- Primary Citation of Related Structures:
2G60 - PubMed Abstract:
The inherent difficulties of stabilizing detergent-solubilized integral membrane proteins for biophysical or structural analysis demand the development of new methodologies to improve success rates. One proven strategy is the use of antibody fragments to increase the ;soluble' portion of any membrane protein, but this approach is limited by the difficulties and expense associated with producing monoclonal antibodies to an appropriate exposed epitope on the target protein. Here, the stabilization of a detergent-solubilized K(+) channel protein, KvPae, by engineering a FLAG-binding epitope into a known loop region of the protein and creating a complex with Fab fragments from commercially available anti-FLAG M2 monoclonal antibodies is reported. Although well diffracting crystals of the complex have not yet been obtained, during the course of crystallization trials the structure of the anti-FLAG M2 Fab domain was solved to 1.86 A resolution. This structure, which should aid future structure-determination efforts using this approach by facilitating molecular-replacement phasing, reveals that the binding pocket appears to be specific only for the first four amino acids of the traditional FLAG epitope, namely DYKD. Thus, the use of antibody fragments for improving the stability of target proteins can be rapidly applied to the study of membrane-protein structure by placing the short DKYD motif within a predicted peripheral loop of that protein and utilizing commercially available anti-FLAG M2 antibody fragments.
Organizational Affiliation:
Structural Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, California 92037, USA.