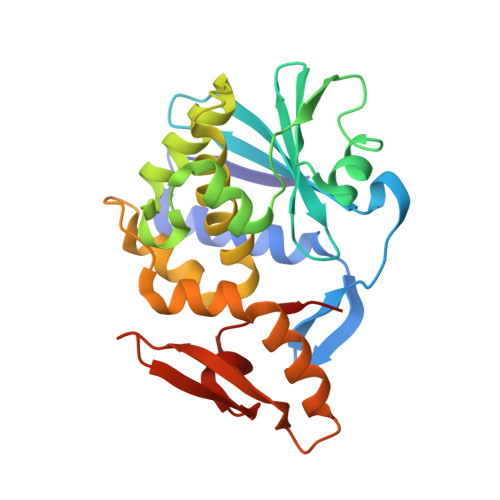
X-ray sequence and crystal structure of luffaculin 1, a novel type 1 ribosome-inactivating protein
Hou, X., Chen, M., Chen, L., Meehan, E.J., Xie, J., Huang, M.(2007) BMC Struct Biol 7: 29-29
- PubMed: 17470286
- DOI: https://doi.org/10.1186/1472-6807-7-29
- Primary Citation of Related Structures:
2OQA - PubMed Abstract:
Protein sequence can be obtained through Edman degradation, mass spectrometry, or cDNA sequencing. High resolution X-ray crystallography can also be used to derive protein sequence information, but faces the difficulty in distinguishing the Asp/Asn, Glu/Gln, and Val/Thr pairs. Luffaculin 1 is a new type 1 ribosome-inactivating protein (RIP) isolated from the seeds of Luffa acutangula. Besides rRNA N-glycosidase activity, luffaculin 1 also demonstrates activities including inhibiting tumor cells' proliferation and inducing tumor cells' differentiation. The crystal structure of luffaculin 1 was determined at 1.4 A resolution. Its amino-acid sequence was derived from this high resolution structure using the following criteria: 1) high resolution electron density; 2) comparison of electron density between two molecules that exist in the same crystal; 3) evaluation of the chemical environment of residues to break down the sequence assignment ambiguity in residue pairs Glu/Gln, Asp/Asn, and Val/Thr; 4) comparison with sequences of the homologous proteins. Using the criteria 1 and 2, 66% of the residues can be assigned. By incorporating with criterion 3, 86% of the residues were assigned, suggesting the effectiveness of chemical environment evaluation in breaking down residue ambiguity. In total, 94% of the luffaculin 1 sequence was assigned with high confidence using this improved X-ray sequencing strategy. Two N-acetylglucosamine moieties, linked respectively to the residues Asn77 and Asn84, can be identified in the structure. Residues Tyr70, Tyr110, Glu159 and Arg162 define the active site of luffaculin 1 as an RNA N-glycosidase. X-ray sequencing method can be effective to derive sequence information of proteins. The evaluation of the chemical environment of residues is a useful method to break down the assignment ambiguity in Glu/Gln, Asp/Asn, and Val/Thr pairs. The sequence and the crystal structure confirm that luffaculin 1 is a new type 1 RIP.
- State Key Laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, The Chinese Academy of Sciences, 155 Yang Qiao Xi Lu, Fuzhou, Fujian, China. houxiaomin@fjirsm.ac.cn <houxiaomin@fjirsm.ac.cn>
Organizational Affiliation: