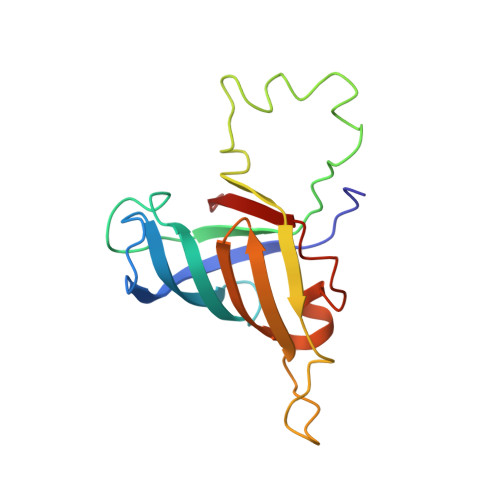
Structural, biochemical, and dynamic characterizations of the hRPB8 subunit of human RNA polymerases
Kang, X., Hu, Y., Li, Y., Guo, X., Jiang, X., Lai, L., Xia, B., Jin, C.(2006) J Biol Chem 281: 18216-18226
- PubMed: 16632472
- DOI: https://doi.org/10.1074/jbc.M513241200
- Primary Citation of Related Structures:
2F3I - PubMed Abstract:
The RPB8 subunit is present in all three types of eukaryotic RNA polymerases and is highly conserved during evolution. It is an essential subunit required for the transcription of nuclear genes, but the detailed mechanism including its interactions with different subunits and oligonucleotides remains largely unclear. Herein, we report the three-dimensional structure of human RPB8 (hRPB8) at high resolution determined by NMR spectroscopy. The protein fold comprises an eight-stranded beta-barrel, six short helices, and a large unstructured Omega-loop. The overall structure of hRPB8 is similar to that of yRPB8 from Saccharomyces cerevisiae and belongs to the oligonucleotide/oligosaccharide-binding fold. However, several features of the tertiary structures are notably different between the two proteins. In particular, hRPB8 has a more clustered positively charged binding interface with the largest subunit RPB1 of the RNA polymerases. We employed biochemical methods to detect its interactions with different single-stranded DNA sequences. In addition, single-stranded DNA titration experiments were performed to identify the residues involved in nonspecific binding with different DNA sequences. Furthermore, we characterized the millisecond time scale conformational flexibility of hRPB8 upon its binding to single-stranded DNA. The current results demonstrate that hRPB8 interacts with single-stranded DNA nonspecifically and adopts significant conformational changes, and the hRPB8/single-stranded DNA complex is a fast exchanging system. The solution structure in conjunction with the biochemical and dynamic studies reveal new aspects of this subunit in the molecular assembly and the biological function of the human nuclear RNA polymerases.
Organizational Affiliation:
Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing 100871, China.