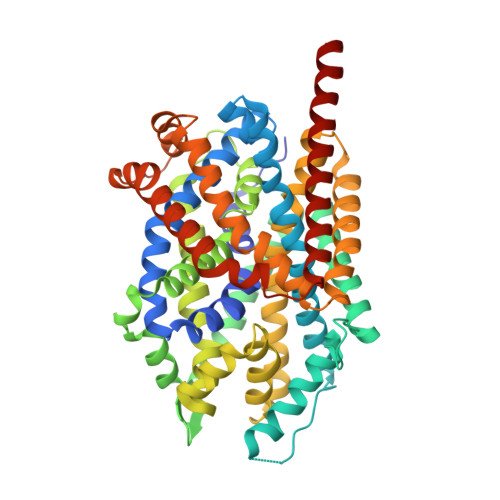
Antidepressant specificity of serotonin transporter suggested by three LeuT-SSRI structures.
Zhou, Z., Zhen, J., Karpowich, N.K., Law, C.J., Reith, M.E., Wang, D.N.(2009) Nat Struct Mol Biol 16: 652-657
- PubMed: 19430461
- DOI: https://doi.org/10.1038/nsmb.1602
- Primary Citation of Related Structures:
3GWU, 3GWV, 3GWW - PubMed Abstract:
Sertraline and fluoxetine are selective serotonin re-uptake inhibitors (SSRIs) that are widely prescribed to treat depression. They exert their effects by inhibiting the presynaptic plasma membrane serotonin transporter (SERT). All SSRIs possess halogen atoms at specific positions, which are key determinants for the drugs' specificity for SERT. For the SERT protein, however, the structural basis of its specificity for SSRIs is poorly understood. Here we report the crystal structures of LeuT, a bacterial SERT homolog, in complex with sertraline, R-fluoxetine or S-fluoxetine. The SSRI halogens all bind to exactly the same pocket within LeuT. Mutation at this halogen-binding pocket (HBP) in SERT markedly reduces the transporter's affinity for SSRIs but not for tricyclic antidepressants. Conversely, when the only nonconserved HBP residue in both norepinephrine and dopamine transporters is mutated into that found in SERT, their affinities for all the three SSRIs increase uniformly. Thus, the specificity of SERT for SSRIs is dependent largely on interaction of the drug halogens with the protein's HBP.
Organizational Affiliation:
Kimmel Center for Biology and Medicine at the Skirball Institute of Biomolecular Medicine, Department of Cell Biology, New York University School of Medicine, New York, New York, USA.