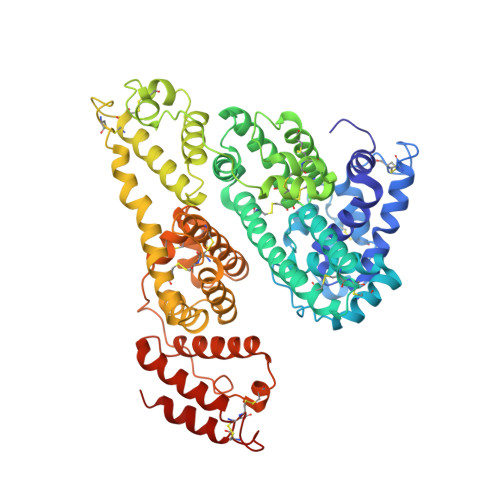
Crystallographic analysis reveals a unique lidocaine binding site on human serum albumin.
Hein, K.L., Kragh-Hansen, U., Morth, J.P., Jeppesen, M.D., Otzen, D., Moller, J.V., Nissen, P.(2010) J Struct Biol
- PubMed: 20347991
- DOI: https://doi.org/10.1016/j.jsb.2010.03.014
- Primary Citation of Related Structures:
3JQZ, 3JRY - PubMed Abstract:
Human serum albumin (HSA), the major protein component in blood plasma and in extravascular spaces, is known to participate in the binding and transport of a variety of endogenous and exogenous organic compounds with anionic or electronegative features. We here report on the 3.3A resolution crystal structure of HSA complexed with the cationic, and widely used, anesthetic lidocaine. We find that lidocaine and HSA co-crystallise as a dimer in the unusual space group I4(1). The dimer consists of one HSA molecule without ligand and one HSA molecule with a single, bound lidocaine. HSA is a heart-shaped protein composed of three homologous helical domains (I-III), which can be subdivided into two subdomains (A and B), and lidocaine binds to a unique site formed by residues from subdomain IB facing the central, interdomain crevice. In the crystal, binding seems to introduce only local conformational changes in the protein. According to intrinsic fluorescence experiments with aqueous HSA binding results in widespread conformational changes involving Trp214 in subdomain IIA. Results obtained with equilibrium dialysis and isothermal titration calorimetry show that lidocaine binding is of a low affinity and occurs at one discrete binding site in accordance with the X-ray data. Another crystal form of ligand-free HSA obtained in the presence of ammonium sulphate was determined at 2.3A resolution revealing a sulphate ion accepting cavity at the surface of subdomain IIIA. The present results contribute to a further characterisation of the exceptional binding properties of HSA.
Organizational Affiliation:
Department of Molecular Biology, University of Aarhus, Gustav Wieds Vej 10C, DK-8000 Arhus C, Denmark.