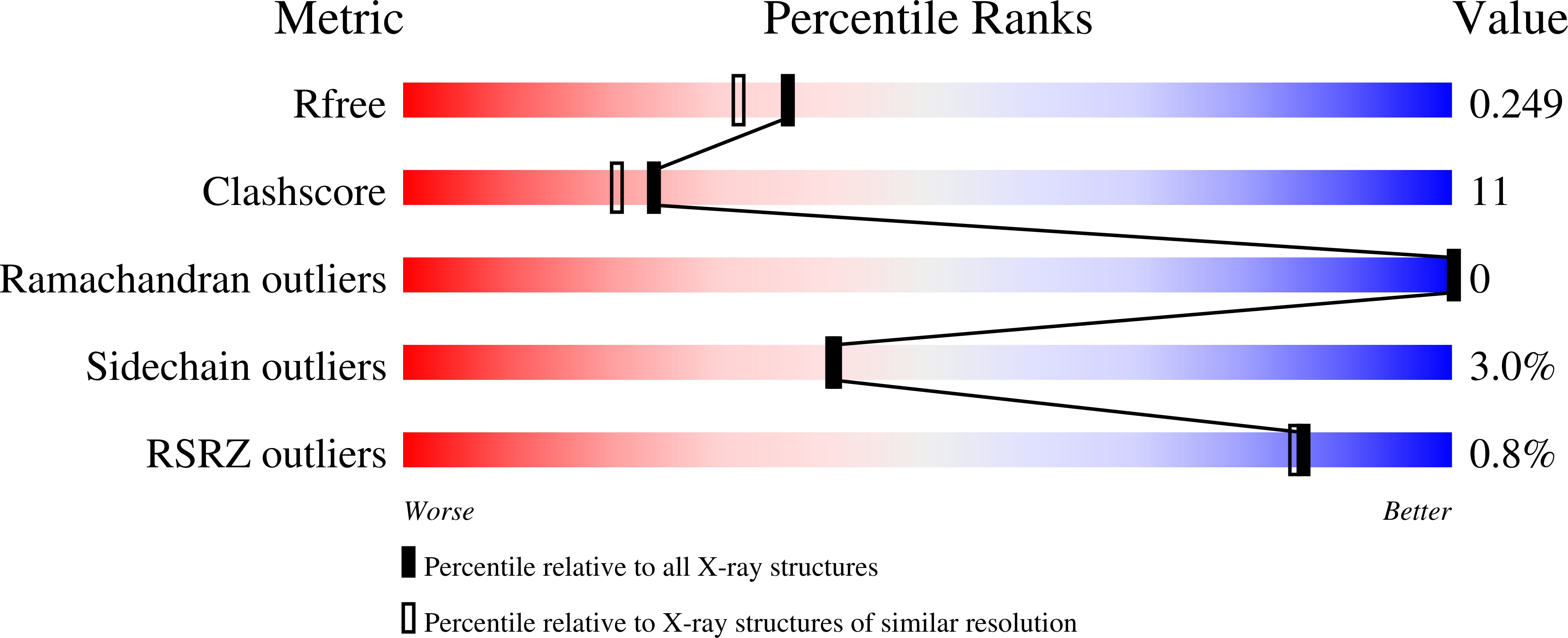
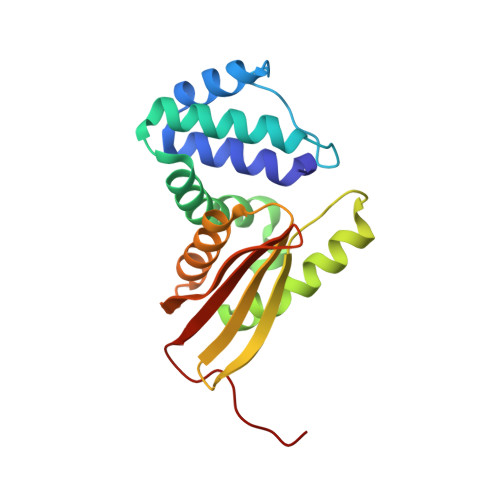
Structural insights into the molecular mechanism of H-NOX activation.
Olea, C., Herzik, M.A., Kuriyan, J., Marletta, M.A.(2010) Protein Sci 19: 881-887
- PubMed: 20162612
- DOI: https://doi.org/10.1002/pro.357
- Primary Citation of Related Structures:
3LAH, 3LAI - PubMed Abstract:
Nitric oxide (NO) signaling in mammals controls important processes such as smooth muscle relaxation and neurotransmission by the activation of soluble guanylate cyclase (sGC). NO binding to the heme domain of sGC leads to dissociation of the iron-histidine (Fe-His) bond, which is required for enzyme activity. The heme domain of sGC belongs to a larger class of proteins called H-NOX (Heme-Nitric oxide/OXygen) binding domains. Previous crystallographic studies on H-NOX domains demonstrate a correlation between heme bending and protein conformation. It was unclear, however, whether these structural changes were important for signal transduction. Subsequent NMR solution structures of H-NOX proteins show a conformational change upon disconnection of the heme and proximal helix, similar to those observed in the crystallographic studies. The atomic details of these conformational changes, however, are lacking in the NMR structures especially at the heme pocket. Here, a high-resolution crystal structure of an H-NOX mutant mimicking a broken Fe-His bond is reported. This mutant exhibits specific changes in heme conformation and major N-terminal displacements relative to the wild-type H-NOX protein. Fe-His ligation is ubiquitous in all H-NOX domains, and therefore, the heme and protein conformational changes observed in this study are likely to occur throughout the H-NOX family when NO binding leads to rupture of the Fe-His bond.
Organizational Affiliation:
Department of Molecular and Cell Biology, University of California, Berkeley, California 94720, USA.