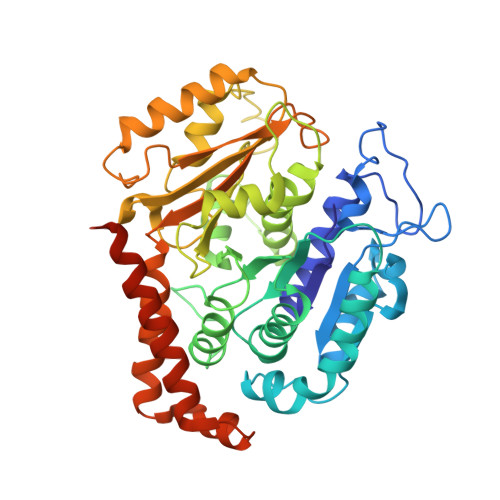
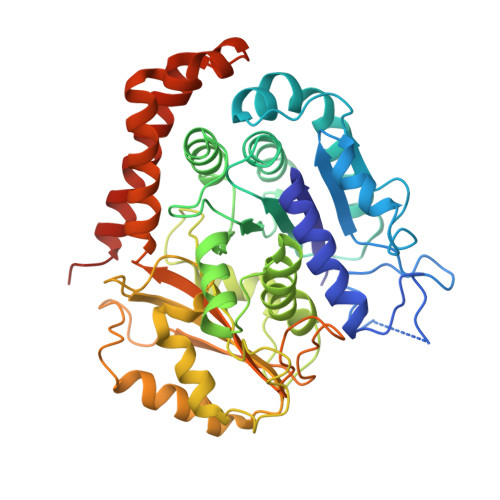
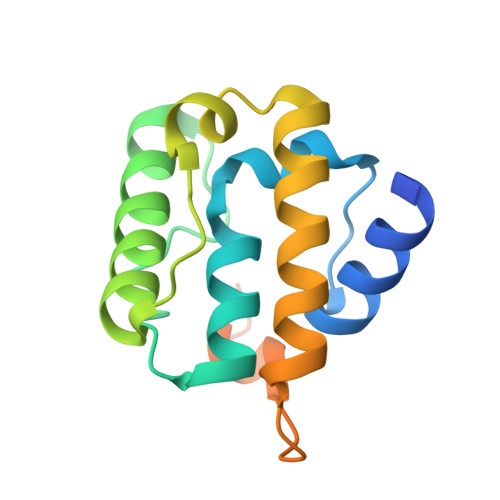
Ebs Recognize a Nucleotide-Dependent Structural CAP at Growing Microtubule Ends.
Maurer, S.P., Fourniol, F.J., Bohner, G., Moores, C.A., Surrey, T.(2012) Cell 149: 371
- PubMed: 22500803
- DOI: https://doi.org/10.1016/j.cell.2012.02.049
- Primary Citation of Related Structures:
4ABO - PubMed Abstract:
Growing microtubule ends serve as transient binding platforms for essential proteins that regulate microtubule dynamics and their interactions with cellular substructures. End-binding proteins (EBs) autonomously recognize an extended region at growing microtubule ends with unknown structural characteristics and then recruit other factors to the dynamic end structure. Using cryo-electron microscopy, subnanometer single-particle reconstruction, and fluorescence imaging, we present a pseudoatomic model of how the calponin homology (CH) domain of the fission yeast EB Mal3 binds to the end regions of growing microtubules. The Mal3 CH domain bridges protofilaments except at the microtubule seam. By binding close to the exchangeable GTP-binding site, the CH domain is ideally positioned to sense the microtubule's nucleotide state. The same microtubule-end region is also a stabilizing structural cap protecting the microtubule from depolymerization. This insight supports a common structural link between two important biological phenomena, microtubule dynamic instability and end tracking.
- Cancer Research UK London Research Institute, Lincoln's Inn Fields Laboratories, 44 Lincoln's Inn Fields, London WC2A 3LY, UK.
Organizational Affiliation: