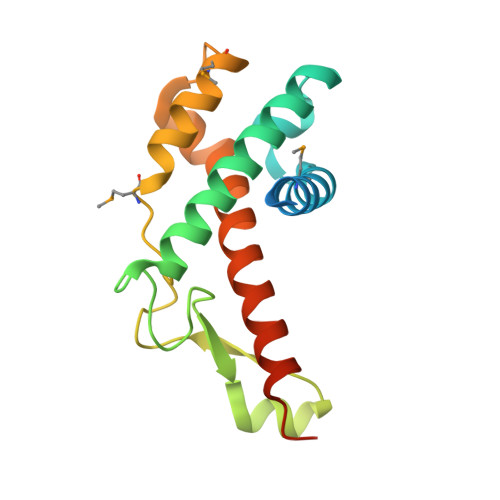
Crystal structure of antitermination protein Q from bacteriophage lambda.
Vorobiev, S., Su, M., Nickels, B., Seetharaman, J., Sahdev, S., Xiao, R., Kogan, S., Maglaqui, M., Wang, D., Everett, J.K., Acton, T.B., Ebright, R.H., Montelione, G.T., Hunt, J., Tong, L.To be published.