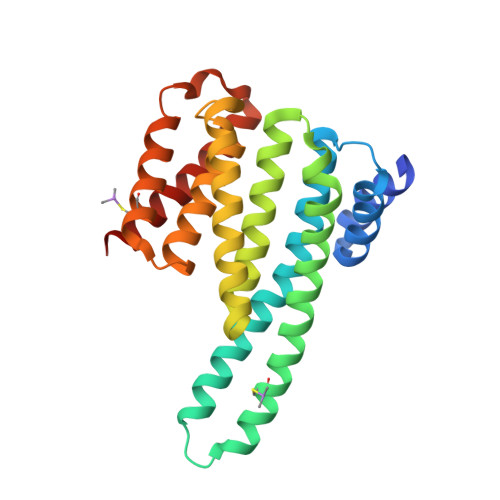
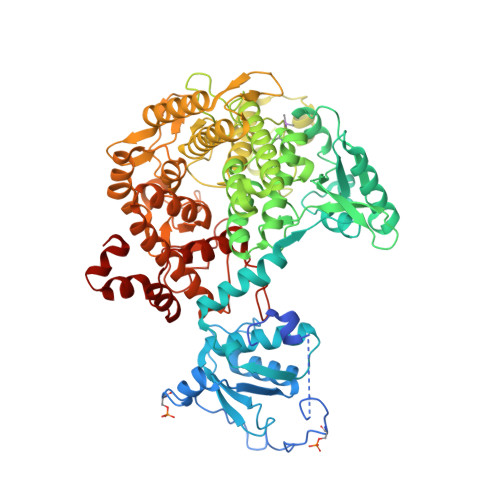
Molecular basis of the 14-3-3 protein-dependent activation of yeast neutral trehalase Nth1.
Alblova, M., Smidova, A., Docekal, V., Vesely, J., Herman, P., Obsilova, V., Obsil, T.(2017) Proc Natl Acad Sci U S A 114: E9811-E9820
- PubMed: 29087344
- DOI: https://doi.org/10.1073/pnas.1714491114
- Primary Citation of Related Structures:
5JTA, 5M4A, 5N6N, 5NIS - PubMed Abstract:
The 14-3-3 proteins, a family of highly conserved scaffolding proteins ubiquitously expressed in all eukaryotic cells, interact with and regulate the function of several hundreds of partner proteins. Yeast neutral trehalases (Nth), enzymes responsible for the hydrolysis of trehalose to glucose, compared with trehalases from other organisms, possess distinct structure and regulation involving phosphorylation at multiple sites followed by binding to the 14-3-3 protein. Here we report the crystal structures of yeast Nth1 and its complex with Bmh1 (yeast 14-3-3 isoform), which, together with mutational and fluorescence studies, indicate that the binding of Nth1 by 14-3-3 triggers Nth1's activity by enabling the proper 3D configuration of Nth1's catalytic and calcium-binding domains relative to each other, thus stabilizing the flexible part of the active site required for catalysis. The presented structure of the Bmh1:Nth1 complex highlights the ability of 14-3-3 to modulate the structure of a multidomain binding partner and to function as an allosteric effector. Furthermore, comparison of the Bmh1:Nth1 complex structure with those of 14-3-3:serotonin N -acetyltransferase and 14-3-3:heat shock protein beta-6 complexes revealed similarities in the 3D structures of bound partner proteins, suggesting the highly conserved nature of 14-3-3 affects the structures of many client proteins.
- Department of Structural Biology of Signaling Proteins, Division Biotechnology and Biomedicine Center of the Academy of Sciences and Charles University in Vestec (BIOCEV), Institute of Physiology, The Czech Academy of Sciences, Prague 14220, Czech Republic.
Organizational Affiliation: