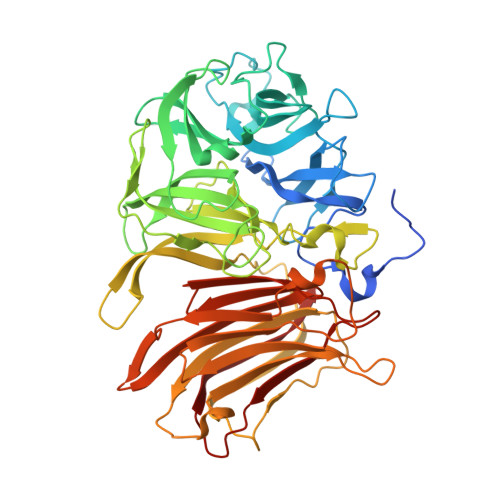
New insights into the molecular mechanism behind mannitol and erythritol fructosylation by beta-fructofuranosidase from Schwanniomyces occidentalis.
Rodrigo-Frutos, D., Jimenez-Ortega, E., Piedrabuena, D., Ramirez-Escudero, M., Miguez, N., Plou, F.J., Sanz-Aparicio, J., Fernandez-Lobato, M.(2021) Sci Rep 11: 7158-7158
- PubMed: 33785821
- DOI: https://doi.org/10.1038/s41598-021-86568-6
- Primary Citation of Related Structures:
6S1T, 6S2B - PubMed Abstract:
The β-fructofuranosidase from Schwanniomyces occidentalis (Ffase) is a useful biotechnological tool for the fructosylation of different acceptors to produce fructooligosaccharides (FOS) and fructo-conjugates. In this work, the structural determinants of Ffase involved in the transfructosylating reaction of the alditols mannitol and erythritol have been studied in detail. Complexes with fructosyl-erythritol or sucrose were analyzed by crystallography and the effect of mutational changes in positions Gln-176, Gln-228, and Asn-254 studied to explore their role in modulating this biocatalytic process. Interestingly, N254T variant enhanced the wild-type protein production of fructosyl-erythritol and FOS by [Formula: see text] 30% and 48%, respectively. Moreover, it produced neokestose, which represented [Formula: see text] 27% of total FOS, and yielded 31.8 g l -1 blastose by using glucose as exclusive fructosyl-acceptor. Noteworthy, N254D and Q176E replacements turned the specificity of Ffase transferase activity towards the synthesis of the fructosylated polyols at the expense of FOS production, but without increasing the total reaction efficiency. The results presented here highlight the relevance of the pair Gln-228/Asn-254 for Ffase donor-sucrose binding and opens new windows of opportunity for optimizing the generation of fructosyl-derivatives by this enzyme enhancing its biotechnological applicability.
- Centro de Biología Molecular Severo Ochoa (CBMSO; UAM-CSIC), Departamento de Biología Molecular, Facultad de Ciencias, Universidad Autónoma de Madrid, Nicolás Cabrera 1, 28049, Madrid, Spain.
Organizational Affiliation: