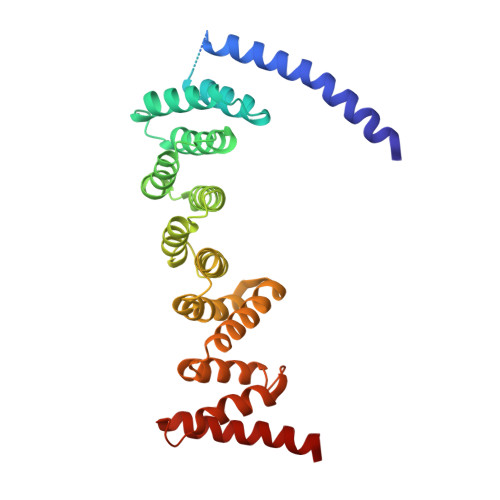
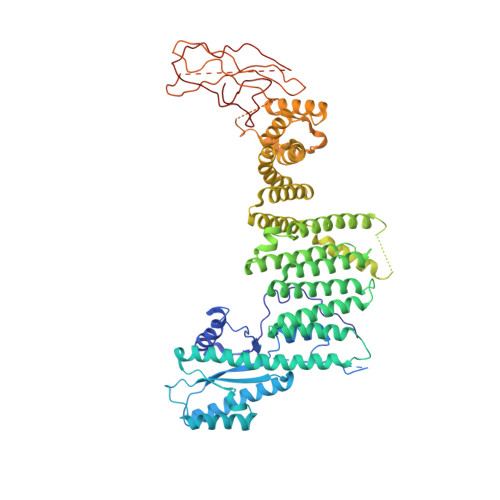
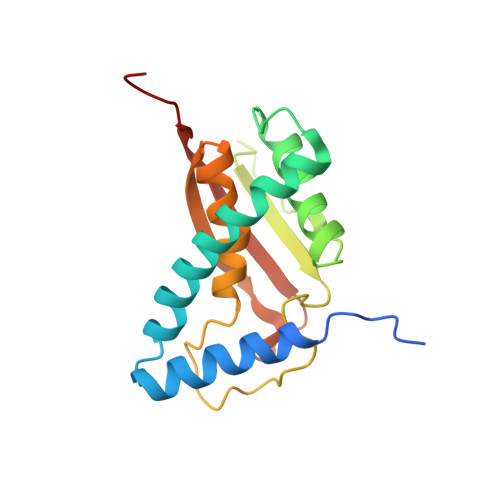
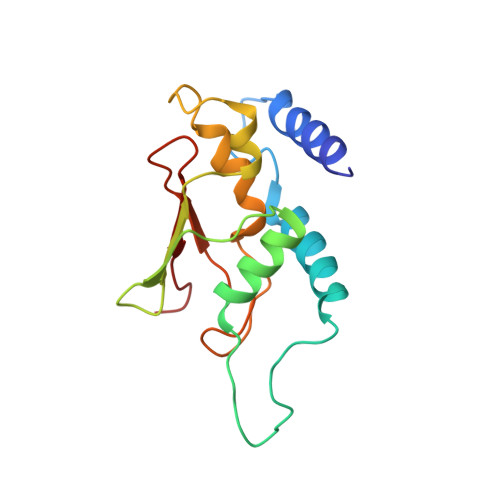
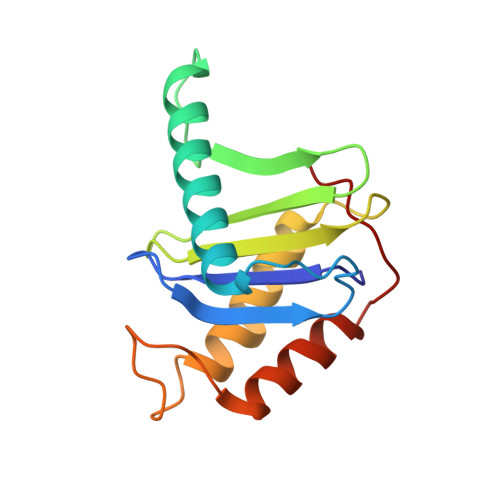
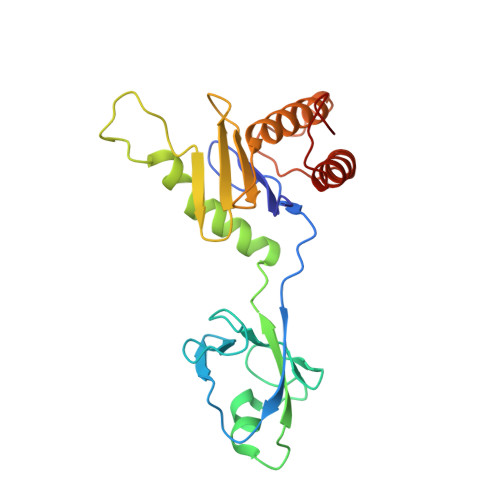
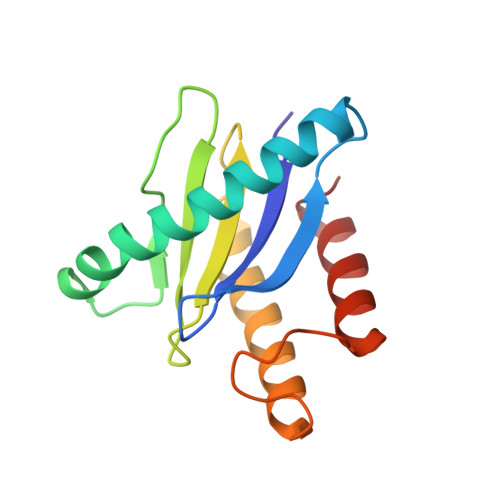
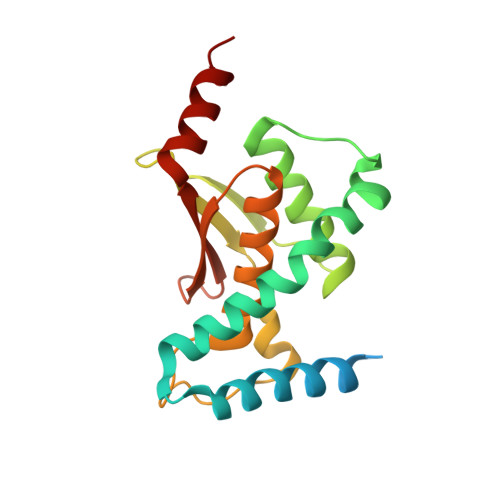
Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Galindo, A., Planelles-Herrero, V.J., Degliesposti, G., Munro, S.(2021) EMBO J 40: e107608-e107608
- PubMed: 34018214
- DOI: https://doi.org/10.15252/embj.2020107608
- Primary Citation of Related Structures:
7B6D, 7B6E, 7B6H, 7B6R, 7B70 - PubMed Abstract:
The TRAPP complexes are nucleotide exchange factors that play essential roles in membrane traffic and autophagy. TRAPPII activates Rab11, and TRAPPIII activates Rab1, with the two complexes sharing a core of small subunits that affect nucleotide exchange but being distinguished by specific large subunits that are essential for activity in vivo. Crystal structures of core subunits have revealed the mechanism of Rab activation, but how the core and the large subunits assemble to form the complexes is unknown. We report a cryo-EM structure of the entire Drosophila TRAPPIII complex. The TRAPPIII-specific subunits TRAPPC8 and TRAPPC11 hold the catalytic core like a pair of tongs, with TRAPPC12 and TRAPPC13 positioned at the joint between them. TRAPPC2 and TRAPPC2L link the core to the two large arms, with the interfaces containing residues affected by disease-causing mutations. The TRAPPC8 arm is positioned such that it would contact Rab1 that is bound to the core, indicating how the arm could determine the specificity of the complex. A lower resolution structure of TRAPPII shows a similar architecture and suggests that the TRAPP complexes evolved from a single ur-TRAPP.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: