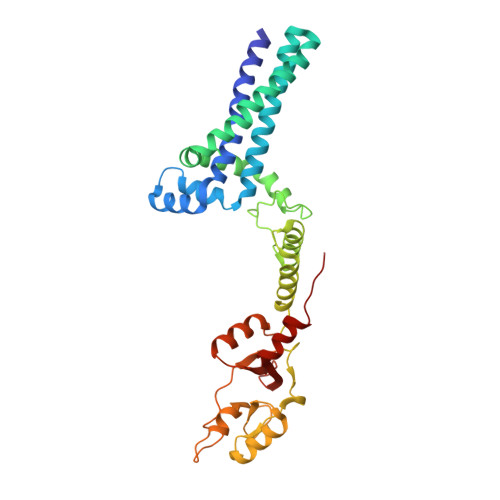
Crystal structure of an archaeal CorB magnesium transporter.
Chen, Y.S., Kozlov, G., Moeller, B.E., Rohaim, A., Fakih, R., Roux, B., Burke, J.E., Gehring, K.(2021) Nat Commun 12: 4028-4028
- PubMed: 34188059
- DOI: https://doi.org/10.1038/s41467-021-24282-7
- Primary Citation of Related Structures:
7M1T, 7M1U, 7MSU - PubMed Abstract:
CNNM/CorB proteins are a broadly conserved family of integral membrane proteins with close to 90,000 protein sequences known. They are associated with Mg 2+ transport but it is not known if they mediate transport themselves or regulate other transporters. Here, we determine the crystal structure of an archaeal CorB protein in two conformations (apo and Mg 2+ -ATP bound). The transmembrane DUF21 domain exists in an inward-facing conformation with a Mg 2+ ion coordinated by a conserved π-helix. In the absence of Mg 2+ -ATP, the CBS-pair domain adopts an elongated dimeric configuration with previously unobserved domain-domain contacts. Hydrogen-deuterium exchange mass spectrometry, analytical ultracentrifugation, and molecular dynamics experiments support a role of the structural rearrangements in mediating Mg 2+ -ATP sensing. Lastly, we use an in vitro, liposome-based assay to demonstrate direct Mg 2+ transport by CorB proteins. These structural and functional insights provide a framework for understanding function of CNNMs in Mg 2+ transport and associated diseases.
- Department of Biochemistry & Centre de Recherche en Biologie Structurale, McGill University, Montréal, QC, Canada.
Organizational Affiliation: