Fragment-Based Discovery of a Novel, Brain Penetrant, Orally Active HDAC2 Inhibitor.
Tamanini, E., Miyamura, S., Buck, I.M., Cons, B.D., Dawson, L., East, C., Futamura, T., Goto, S., Griffiths-Jones, C., Hashimoto, T., Heightman, T.D., Ishikawa, S., Ito, H., Kaneko, Y., Kawato, T., Kondo, K., Kurihara, N., McCarthy, J.M., Mori, Y., Nagase, T., Nakaishi, Y., Reeks, J., Sato, A., Schopf, P., Tai, K., Tamai, T., Tisi, D., Woolford, A.J.(2022) ACS Med Chem Lett 13: 1591-1597
- PubMed: 36262388
- DOI: https://doi.org/10.1021/acsmedchemlett.2c00272
- Primary Citation of Related Structures:
7ZZO, 7ZZP, 7ZZR, 7ZZS, 7ZZT, 7ZZU, 7ZZW, 8A0B - PubMed Abstract:
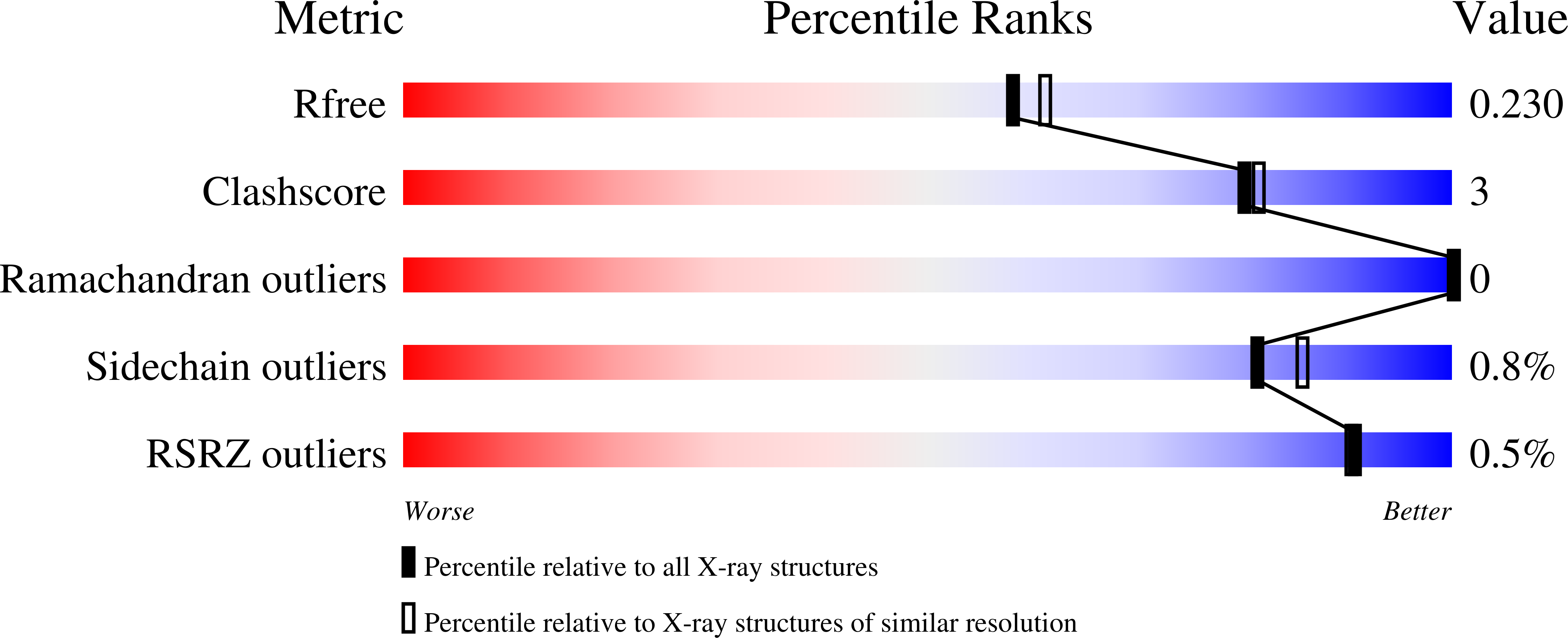
Fragment-based ligand discovery was successfully applied to histone deacetylase HDAC2. In addition to the anticipated hydroxamic acid- and benzamide-based fragment screening hits, a low affinity (∼1 mM) α-amino-amide zinc binding fragment was identified, as well as fragments binding to other regions of the catalytic site. This alternative zinc-binding fragment was further optimized, guided by the structural information from protein-ligand complex X-ray structures, into a sub-μM, brain penetrant, HDAC2 inhibitor ( 17 ) capable of modulating histone acetylation levels in vivo .
Organizational Affiliation:
Astex Pharmaceuticals, 436 Cambridge Science Park, Milton Road, Cambridge CB4 0QA, U.K.