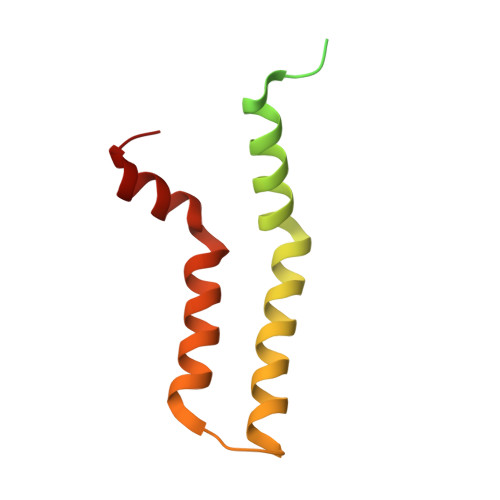
Cryo-EM structure of the Agrobacterium tumefaciens T-pilus reveals the importance of positive charges in the lumen.
Amro, J., Black, C., Jemouai, Z., Rooney, N., Daneault, C., Zeytuni, N., Ruiz, M., Bui, K.H., Baron, C.(2023) Structure 31: 375-384.e4
- PubMed: 36513067
- DOI: https://doi.org/10.1016/j.str.2022.11.007
- Primary Citation of Related Structures:
8CUE, 8CW4 - PubMed Abstract:
Agrobacterium tumefaciens is a natural genetic engineer that transfers DNA into plants, which is the most applied process for generation of genetically modified plants. DNA transfer is mediated by a type IV secretion system in the cell envelope and extracellular T-pili. We here report the cryo-electron microscopic structures of the T-pilus at 3.2-Å resolution and of the plasmid pKM101-determined N-pilus at 3-Å resolution. Both pili contain a main pilus protein (VirB2 in A. tumefaciens, TraM in pKM101) and phospholipids arranged in a five-start helical assembly. They contain positively charged amino acids in the lumen, and the lipids are positively charged in the T-pilus (phosphatidylcholine) conferring overall positive charge. Mutagenesis of the lumen-exposed Arg91 in VirB2 results in protein destabilization and loss of pilus formation. Our results reveal that different phospholipids can be incorporated into type IV secretion pili and that the charge of the lumen may be of functional importance.
- Department of Biochemistry and Molecular Medicine, Faculty of Medicine, Université de Montréal, Québec, QC, Canada.
Organizational Affiliation: