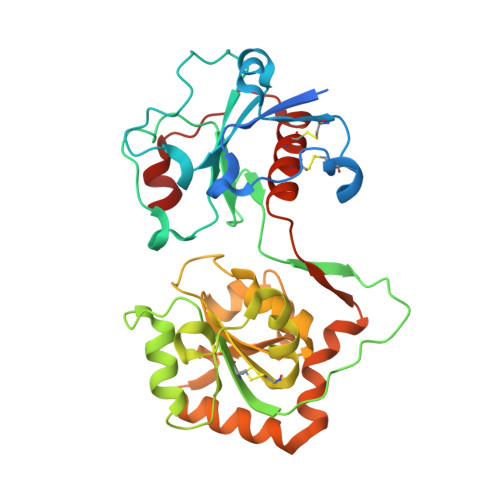
Structural basis for Lewis antigen synthesis by the alpha 1,3-fucosyltransferase FUT9.
Kadirvelraj, R., Boruah, B.M., Wang, S., Chapla, D., Huang, C., Ramiah, A., Hudson, K.L., Prudden, A.R., Boons, G.J., Withers, S.G., Wood, Z.A., Moremen, K.W.(2023) Nat Chem Biol 19: 1022-1030
- PubMed: 37202521
- DOI: https://doi.org/10.1038/s41589-023-01345-y
- Primary Citation of Related Structures:
8D0O, 8D0P, 8D0Q, 8D0R, 8D0S, 8D0U, 8D0W, 8D0X - PubMed Abstract:
Mammalian cell surface and secreted glycoproteins exhibit remarkable glycan structural diversity that contributes to numerous physiological and pathogenic interactions. Terminal glycan structures include Lewis antigens synthesized by a collection of α1,3/4-fucosyltransferases (CAZy GT10 family). At present, the only available crystallographic structure of a GT10 member is that of the Helicobacter pylori α1,3-fucosyltransferase, but mammalian GT10 fucosyltransferases are distinct in sequence and substrate specificity compared with the bacterial enzyme. Here, we determined crystal structures of human FUT9, an α1,3-fucosyltransferase that generates Lewis x and Lewis y antigens, in complex with GDP, acceptor glycans, and as a FUT9-donor analog-acceptor Michaelis complex. The structures reveal substrate specificity determinants and allow prediction of a catalytic model supported by kinetic analyses of numerous active site mutants. Comparisons with other GT10 fucosyltransferases and GT-B fold glycosyltransferases provide evidence for modular evolution of donor- and acceptor-binding sites and specificity for Lewis antigen synthesis among mammalian GT10 fucosyltransferases.
- Department of Biochemistry & Molecular Biology, University of Georgia, Athens, GA, USA.
Organizational Affiliation: