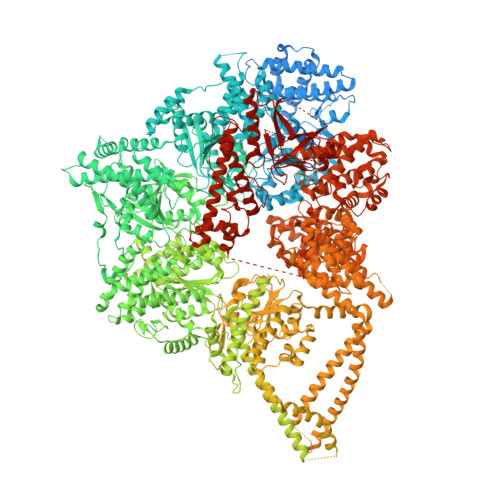
Structures of human dynein in complex with the lissencephaly 1 protein, LIS1.
Reimer, J.M., DeSantis, M.E., Reck-Peterson, S.L., Leschziner, A.E.(2023) Elife 12
- PubMed: 36692009
- DOI: https://doi.org/10.7554/eLife.84302
- Primary Citation of Related Structures:
8DYU, 8DYV - PubMed Abstract:
The lissencephaly 1 protein, LIS1, is mutated in type-1 lissencephaly and is a key regulator of cytoplasmic dynein-1. At a molecular level, current models propose that LIS1 activates dynein by relieving its autoinhibited form. Previously we reported a 3.1 Å structure of yeast dynein bound to Pac1, the yeast homologue of LIS1, which revealed the details of their interactions (Gillies et al., 2022). Based on this structure, we made mutations that disrupted these interactions and showed that they were required for dynein's function in vivo in yeast. We also used our yeast dynein-Pac1 structure to design mutations in human dynein to probe the role of LIS1 in promoting the assembly of active dynein complexes. These mutations had relatively mild effects on dynein activation, suggesting that there may be differences in how dynein and Pac1/LIS1 interact between yeast and humans. Here, we report cryo-EM structures of human dynein-LIS1 complexes. Our new structures reveal the differences between the yeast and human systems, provide a blueprint to disrupt the human dynein-LIS1 interactions more accurately, and map type-1 lissencephaly disease mutations, as well as mutations in dynein linked to malformations of cortical development/intellectual disability, in the context of the dynein-LIS1 complex.
Organizational Affiliation:
Department of Cellular and Molecular Medicine, University of California, San Diego, San Diego, United States.