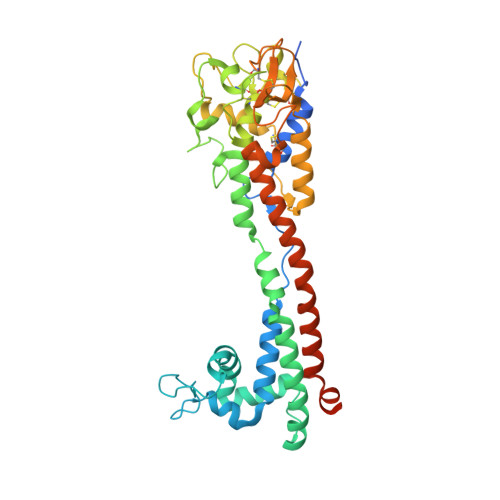
A structural classification of the variant surface glycoproteins of the African trypanosomey.
Dakovic, S., Zeelen, J.P., Gkeka, A., Chandra, M., van Straaten, M., Foti, K., Zhong, J., Vlachou, E.P., Aresta-Branco, F., Verdi, J.P., Papavasiliou, F.N., Stebbins, C.E.(2023) PLoS Negl Trop Dis 17: e0011621-e0011621
- PubMed: 37656766
- DOI: https://doi.org/10.1371/journal.pntd.0011621
- Primary Citation of Related Structures:
8OK4, 8OK5, 8OK6, 8OK7, 8OK8, 8ONH, 8Q0E, 8Q0P - PubMed Abstract:
Long-term immune evasion by the African trypanosome is achieved through repetitive cycles of surface protein replacement with antigenically distinct versions of the dense Variant Surface Glycoprotein (VSG) coat. Thousands of VSG genes and pseudo-genes exist in the parasite genome that, together with genetic recombination mechanisms, allow for essentially unlimited immune escape from the adaptive immune system of the host. The diversity space of the "VSGnome" at the protein level was thought to be limited to a few related folds whose structures were determined more than 30 years ago. However, recent progress has shown that the VSGs possess significantly more architectural variation than had been appreciated. Here we combine experimental X-ray crystallography (presenting structures of N-terminal domains of coat proteins VSG11, VSG21, VSG545, VSG558, and VSG615) with deep-learning prediction using Alphafold to produce models of hundreds of VSG proteins. We classify the VSGnome into groups based on protein architecture and oligomerization state, contextualize recent bioinformatics clustering schemes, and extensively map VSG-diversity space. We demonstrate that in addition to the structural variability and post-translational modifications observed thus far, VSGs are also characterized by variations in oligomerization state and possess inherent flexibility and alternative conformations, lending additional variability to what is exposed to the immune system. Finally, these additional experimental structures and the hundreds of Alphafold predictions confirm that the molecular surfaces of the VSGs remain distinct from variant to variant, supporting the hypothesis that protein surface diversity is central to the process of antigenic variation used by this organism during infection.
- Division of Structural Biology of Infection and Immunity, German Cancer Research Center, Heidelberg, Germany.
Organizational Affiliation: