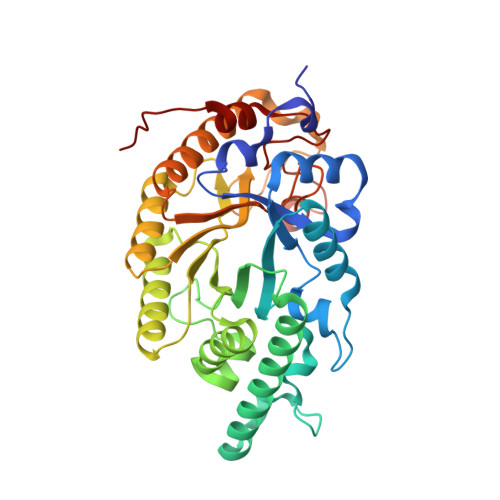
Activity-stability trade-off observed in variants at position 315 of the GH10 xylanase XynR.
Nakamura, T., Takita, T., Kuwata, K., Mizutani, K., Mikami, B., Nakamura, S., Yasukawa, K.(2024) Sci Rep 14: 7767-7767
- PubMed: 38565938
- DOI: https://doi.org/10.1038/s41598-024-57819-z
- Primary Citation of Related Structures:
8XY0, 8Y1M - PubMed Abstract:
XynR is a thermostable alkaline GH10 xylanase, for which we have previously examined the effects of saturation mutagenesis at position 315 on enzyme alkaliphily, and found that at pH 10, the activities of variants could be ordered as follows: T315Q > T315S = T315N > T315H = wild-type XynR (WT) > 15 other variants. In this study, we sought to elucidate the mechanisms underlying the variable activity of these different variants. Crystallographic analysis revealed that the Ca 2+ ion near position 315 in WT was absent in the T315Q variant. We accordingly hypothesized that the enhancement of alkaliphily in T315Q, and probably also in the T315H, T315N, and T315S variants, could be ascribed to an activity-stability trade-off associated with a reduction in stability due to the lack of this Ca 2+ ion. Consistent with expectations, the alkaline resistance of T315H, T315N, T315Q, and T315S, evaluated through the pH-dependence of stability at 0 mM CaCl 2 under alkaline conditions, was found to be lower than that of WT: the residual activity at pH 11 of WT was 78% while those of T315H, T315N, T315Q, and T315S were 0, 9, 0, and 43%, respectively. In addition, the thermostabilities of these four variants, as assessed using the denaturing temperatures (T m ) at 0 mM CaCl 2 based on ellipticity at 222 nm in circular dichroism measurements, were lower than that of WT by 2-8 °C. Furthermore, the T m values of WT and variants at 5 mM CaCl 2 were higher than those at 0 mM CaCl 2 by 6-11 °C. Collectively, our findings in this study indicate that mutation of the T residue at position 315 of XynR to H, N, Q, and S causes an increase in the alkaliphily of this enzyme, thereby reducing its stability.
- Division of Food Science and Biotechnology, Graduate School of Agriculture, Kyoto University, Sakyo-ku, Kyoto, 606-8502, Japan.
Organizational Affiliation: