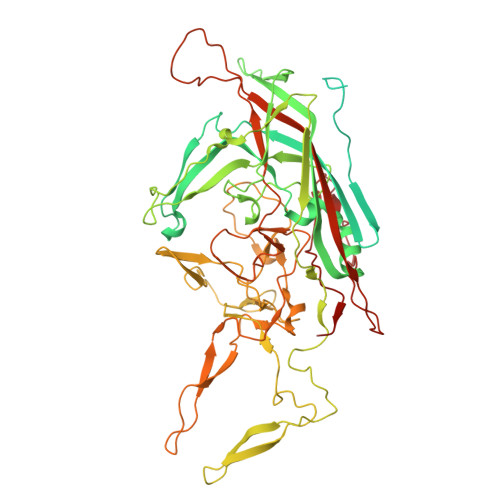
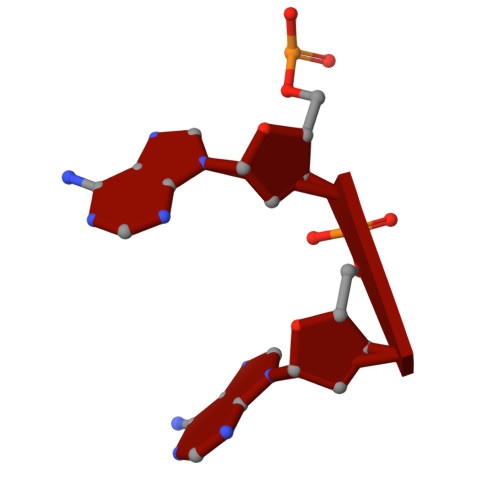
Biophysical and structural insights into AAV genome ejection.
Gliwa, K., Hull, J., Kansol, A., Zembruski, V., Lakshmanan, R., Mietzsch, M., Chipman, P., Bennett, A., McKenna, R.(2025) J Virol 99: e0089924-e0089924
- PubMed: 39907279
- DOI: https://doi.org/10.1128/jvi.00899-24
- Primary Citation of Related Structures:
9DC7, 9DCB, 9DCC - PubMed Abstract:
Recombinant adeno-associated virus (rAAV) is comprised of non-enveloped capsids that can package a therapeutic transgene and are currently being developed and utilized as gene therapy vectors. The therapeutic efficiency of rAAV is dependent on successful cytoplasmic trafficking and transgene delivery to the nucleus. It is hypothesized that an increased understanding of the effects of the cellular environment and biophysical properties of the capsid as it traffics to the nucleus could provide insight to improve vector efficiency. The AAV capsid is exposed to increasing [H + ] during endo-lysosomal trafficking. Exposure to low pH facilitates the externalization of the viral protein 1 unique region (VP1u). This VP1u contains a phospholipase A2 domain required for endosomal escape and nuclear localization signals that facilitate nuclear targeting and entry. The viral genome is released either after total capsid disassembly or via a concerted DNA ejection mechanism in the nucleus. This study presents the characterization of genome ejection (GE) for two diverse serotypes, AAV2 and AAV5, using temperature. The temperature required to disassemble the virus capsid (T M ) is significantly higher than the temperature required to expose the transgene (T E ) for both serotypes. This was verified by quantitative PCR (qPCR) and transmission electron microscopy. Additionally, the absence of VP1/VP2 in the capsids and a decrease in pH increase the temperature of GE. Furthermore, cryo-electron microscopy structures of the AAV5 capsid pre- and post-GE reveal dynamics at the twofold, threefold, and fivefold regions of the capsid interior consistent with a concerted egress of the viral genome.IMPORTANCEThe development of recombinant adeno-associated virus (rAAV) capsids has grown rapidly in recent years, with five of the eight established therapeutics gaining approval in the past 2 years alone. Clinical progression with AAV2 and AAV5 represents a growing need to further characterize the molecular biology of these viruses. The goal of AAV-based gene therapy is to treat monogenic disorders with a vector-delivered transgene to provide wild-type protein function. A better understanding of the dynamics and conditions enabling transgene release may improve therapeutic efficiency. In addition to their clinical importance, AAV2 and 5 were chosen in this study for their diverse antigenic and biophysical properties compared to more closely related serotypes. Characterization of a shared genome ejection process may imply a conserved mechanism for all rAAV therapies.
- Department of Biochemistry and Molecular Biology, University of Florida, Gainesville, Florida, USA.
Organizational Affiliation: