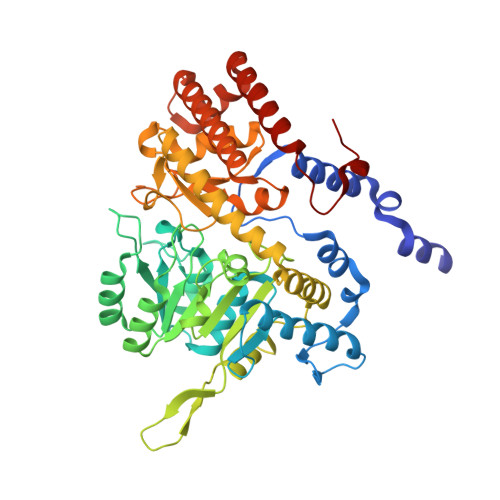
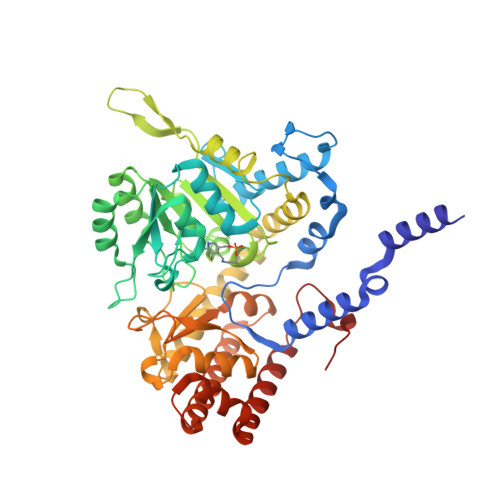
Structural basis of methotrexate and pemetrexed action on serine hydroxymethyltransferases revealed using plant models.
Ruszkowski, M., Sekula, B., Ruszkowska, A., Contestabile, R., Nogues, I., Angelaccio, S., Szczepaniak, A., Dauter, Z.(2019) Sci Rep 9: 19614-19614
- PubMed: 31873125
- DOI: https://doi.org/10.1038/s41598-019-56043-4
- Primary Citation of Related Structures:
6SMN, 6SMR, 6SMW - PubMed Abstract:
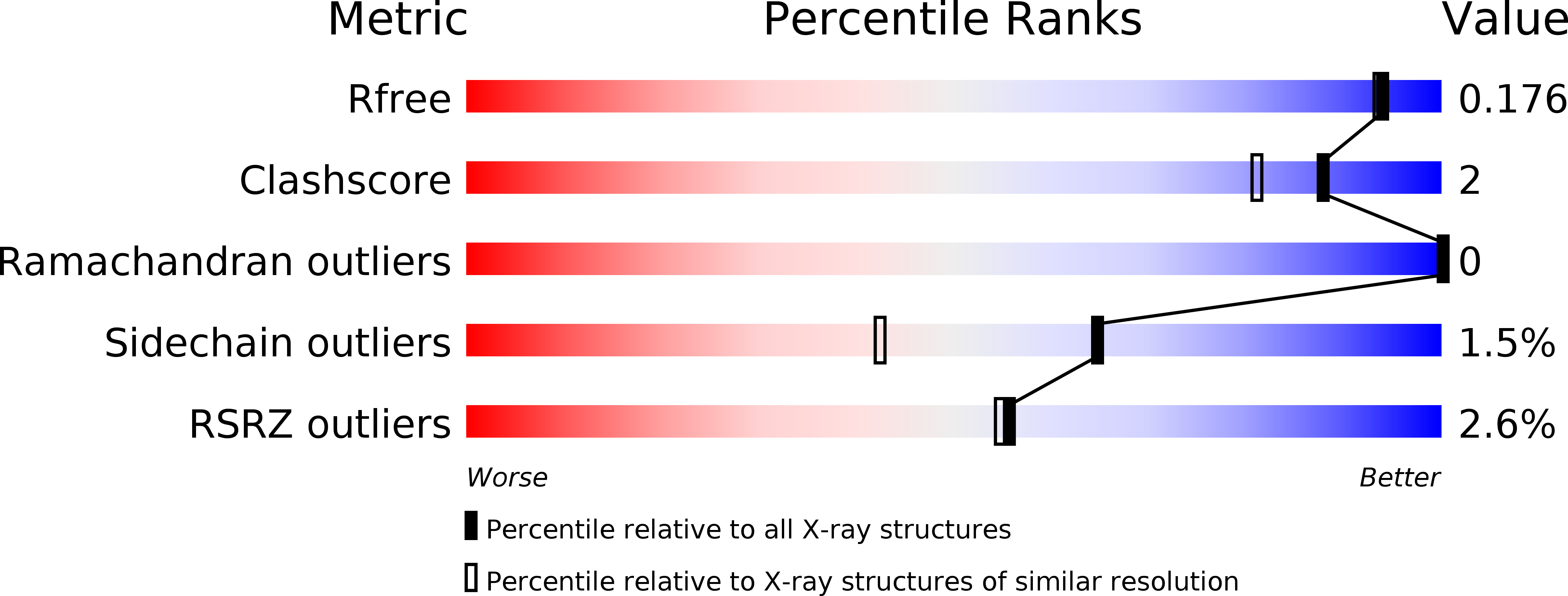
Serine hydroxymethyltransferases (SHMTs) reversibly transform serine into glycine in a reaction accompanied with conversion of tetrahydrofolate (THF) into 5,10-methylene-THF (5,10-meTHF). In vivo, 5,10-meTHF is the main carrier of one-carbon (1C) units, which are utilized for nucleotide biosynthesis and other processes crucial for every living cell, but hyperactivated in overproliferating cells (e.g. cancer tissues). SHMTs are emerging as a promising target for development of new drugs because it appears possible to inhibit growth of cancer cells by cutting off the supply of 5,10-meTHF. Methotrexate (MTX) and pemetrexed (PTX) are two examples of antifolates that have cured many patients over the years but target different enzymes from the folate cycle (mainly dihydrofolate reductase and thymidylate synthase, respectively). Here we show crystal structures of MTX and PTX bound to plant SHMT isozymes from cytosol and mitochondria-human isozymes exist in the same subcellular compartments. We verify inhibition of the studied isozymes by a thorough kinetic analysis. We propose to further exploit antifolate scaffold in development of SHMT inhibitors because it seems likely that especially polyglutamylated PTX inhibits SHMTs in vivo. Structure-based optimization is expected to yield novel antifolates that could potentially be used as chemotherapeutics.
Organizational Affiliation:
Synchrotron Radiation Research Section of MCL, National Cancer Institute, Argonne, IL, USA. mruszkowski@ibch.poznan.pl.