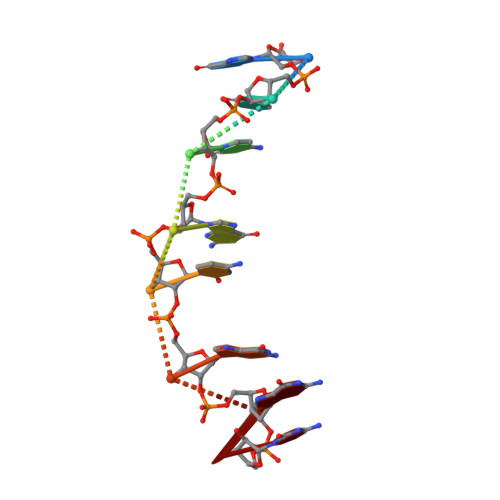
Solution structure and thermodynamics of 2',5' RNA intercalation.
Horowitz, E.D., Lilavivat, S., Holladay, B.W., Germann, M.W., Hud, N.V.(2009) J Am Chem Soc 131: 5831-5838
- PubMed: 19309071
- DOI: https://doi.org/10.1021/ja810068e
- Primary Citation of Related Structures:
2KD4 - PubMed Abstract:
As a means to explore the influence of the nucleic acid backbone on the intercalative binding of ligands to DNA and RNA, we have determined the solution structure of a proflavine-bound 2',5'-linked octamer duplex with the sequence GCCGCGGC. This structure represents the first NMR structure of an intercalated RNA duplex, of either backbone structural isomer. By comparison with X-ray crystal structures, we have identified similarities and differences between intercalated 3',5' and 2',5'-linked RNA duplexes. First, the two forms of RNA have different sugar pucker geometries at the intercalated nucleotide steps, yet have the same interphosphate distances. Second, as in intercalated 3',5' RNA, the phosphate backbone angle zeta at the 2',5' RNA intercalation site prefers to be in the trans conformation, whereas unintercalated 2',5' and 3',5' RNA prefer the -gauche conformation. These observations provide new insights regarding the transitions required for intercalation of a phosphodiester-ribose backbone and suggest a possible contribution of the backbone to the origin of the nearest-neighbor exclusion principle. Thermodynamic studies presented for intercalation of both structural RNA isomers also reveal a surprising sensitivity of intercalator binding enthalpy and entropy to the details of RNA backbone structure.
- Parker H. Petit Institute of Bioengineering and Bioscience, Georgia Institute of Technology, School of Chemistry and Biochemistry, Atlanta, Georgia 30332-0400, USA.
Organizational Affiliation: