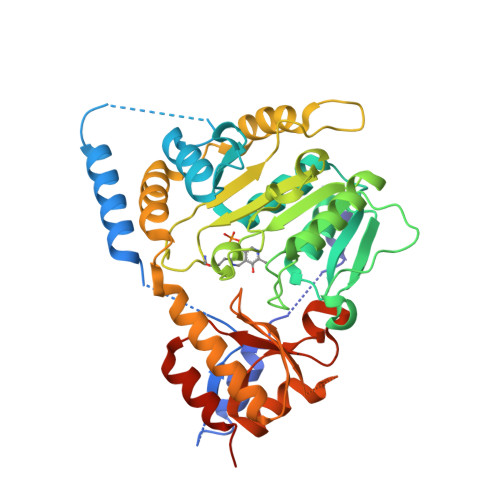
The Structure of ll-Diaminopimelate Aminotransferase from Chlamydia trachomatis: Implications for Its Broad Substrate Specificity.
Watanabe, N., Clay, M.D., van Belkum, M.J., Fan, C., Vederas, J.C., James, M.N.(2011) J Mol Biology 411: 649-660
- PubMed: 21722650
- DOI: https://doi.org/10.1016/j.jmb.2011.06.023
- Primary Citation of Related Structures:
3ASA, 3ASB - PubMed Abstract:
We have previously reported the structures of the native holo and substrate-bound forms of LL-diaminopimelate aminotransferase from Arabidopsis thaliana (AtDAP-AT). Here, we report the crystal and molecular structures of the LL-diaminopimelate aminotransferase from Chlamydia trachomatis (CtDAP-AT) in the apo-form and the pyridoxal-5'-phosphate-bound form. The molecular structure of CtDAP-AT shows that its overall fold is essentially identical with that of AtDAP-AT except that CtDAP-AT adopts an "open" conformation as opposed to the "closed" conformation of AtDAP-AT. Although AtDAP-AT and CtDAP-AT are approximately 40% identical in their primary sequence, they have major differences in their substrate specificities; AtDAP-AT is highly specific for LL-DAP, whereas CtDAP-AT accepts a wider range of substrates. Since all of the residues involved in substrate recognition are highly conserved between AtDAP-AT and CtDAP-AT, we propose that differences in flexibility of the loops lining the active-site region between the two enzymes likely account for the differences in substrate specificity.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada.
Organizational Affiliation: