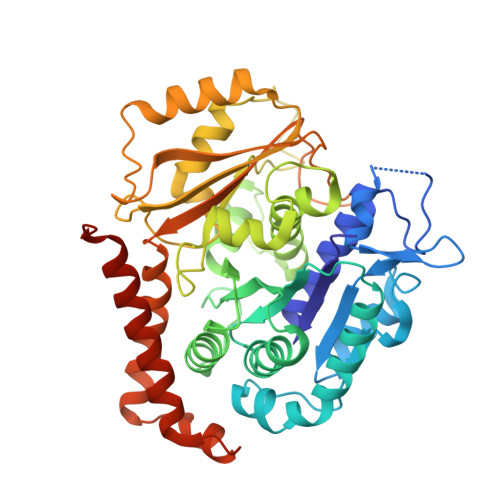
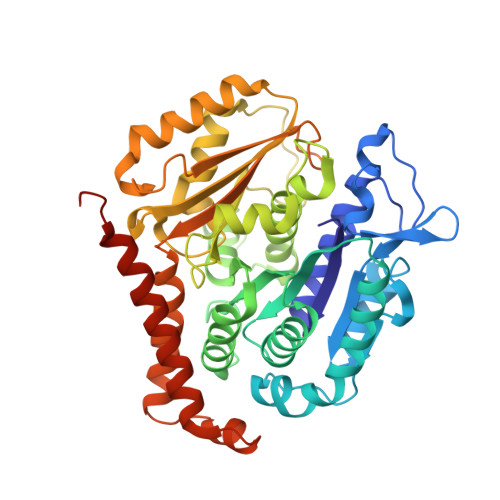
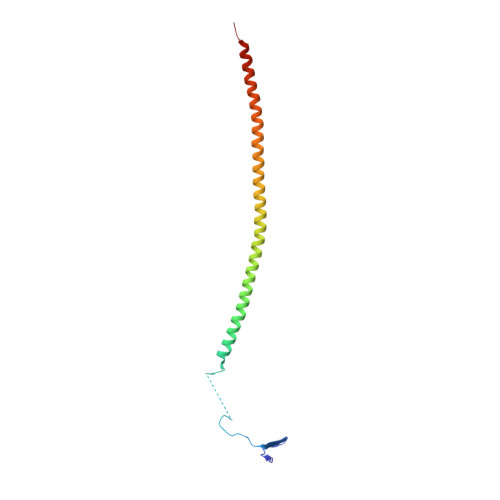
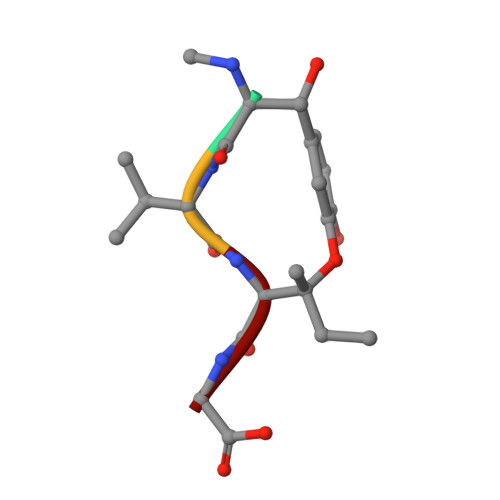
Structural plasticity of tubulin assembly probed by vinca-domain ligands.
Ranaivoson, F.M., Gigant, B., Berritt, S., Joullie, M., Knossow, M.(2012) Acta Crystallogr D Biol Crystallogr 68: 927-934
- PubMed: 22868758
- DOI: https://doi.org/10.1107/S0907444912017143
- Primary Citation of Related Structures:
3UT5, 4EB6 - PubMed Abstract:
Vinca-domain ligands are compounds that bind to tubulin at its inter-heterodimeric interface and favour heterogeneous protofilament-like assemblies, giving rise to helices and rings. This is the basis for their inhibition of microtubule assembly, for their antimitotic activities and for their use in anticancer chemotherapy. Ustiloxins are vinca-domain ligands with a well established total synthesis. A 2.7 Å resolution structure of ustiloxin D bound to the vinca domain embedded in the complex of two tubulins with the stathmin-like domain of RB3 (T(2)R) has been determined. This finding precisely defines the interactions of ustiloxins with tubulin and, taken together with structures of other vinca-ligand complexes, allows structure-based suggestions to be made for improved activity. These comparisons also provide a rationale for the large-scale polymorphism of the protofilament-like assemblies mediated by vinca-domain ligands based on local differences in their interactions with the two tubulin heterodimers constituting their binding site.
- Laboratoire d'Enzymologie et Biochimie Structurales, Centre de Recherche de Gif, CNRS, 91198 Gif-sur-Yvette, France.
Organizational Affiliation: